Short Answer: Yes — and this may be the most compelling near-term application of airborne eDNA technology. Multiple studies have demonstrated detection of plant pathogens (particularly fungal diseases affecting wheat, barley and other cereals) from air samples, in some cases before visible symptoms appear. The combination of molecular sensitivity, non-targeted surveillance capability, and the existing infrastructure of agricultural pollen monitoring stations (already deployed at field-scale across major arable regions) makes this one of the most tractable near-term deployment scenarios.
1. The crop protection problem
Crop diseases and agricultural pests destroy up to 40 per cent of global crop production annually, with plant diseases alone costing the global economy an estimated $220 billion per year (FAO). In temperate Europe, fungal diseases of cereals — rust, powdery mildew, Fusarium, Septoria — are managed primarily through prophylactic fungicide applications timed to risk calendars rather than actual detected presence. This results in both over-application (high cost, environmental impact) and under-application (missed treatment windows when disease arrives unexpectedly).
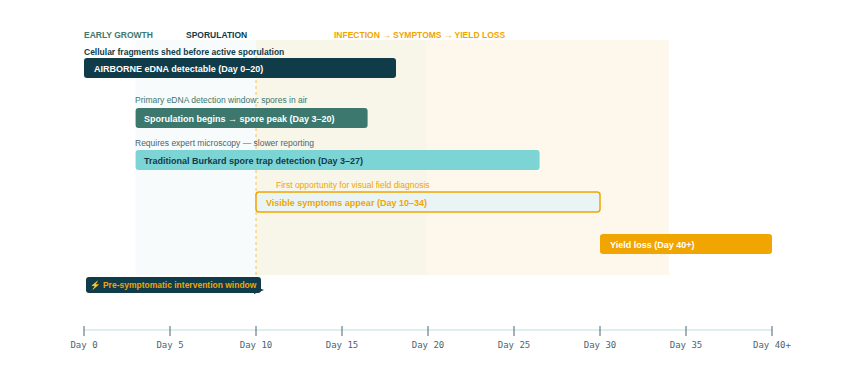
Surveillance-led, responsive management — treating only when and where disease is actually detected — requires a monitoring system that detects pathogen spores in the air before they cause visible symptoms. This is exactly what the Burkard spore trap network has attempted to provide for decades for several priority pathogens. But Burkard traps require microscopic identification by expert analysts, which limits the number of species that can be monitored and the speed of reporting.
Airborne eDNA metabarcoding and shotgun sequencing transform this limitation: a single air sample can be screened for all known crop pathogens and pests simultaneously, with automated bioinformatic analysis, at a fraction of the cost of expert microscopy.
A secondary and currently under-explored detection opportunity are emerging fungal fruiting bodies and mycelial structures that shed DNA-bearing cellular material that could be detectable via airborne eDNA before active sporulation begins. Analogous to the detection of plants from the air via non-pollen leaf fragments outside the pollen season, this would extend the detection window even earlier in the disease cycle. No published study has yet demonstrated pre-sporulation crop pathogen detection via airborne eDNA, but the principle is scientifically plausible and worth investigating.
2. Demonstrated airborne eDNA detection of crop pathogens
2.1 Mikko et al. (2025): surveillance of crop pathogens and pests
Mikko et al. (2025) is the most directly relevant study. Working from the same Swedish research group as Sullivan et al. (2025), they used shotgun sequencing of airborne eDNA to detect crop pathogens and pests at an agricultural monitoring site in Sweden. They demonstrated:
- Detection of multiple fungal wheat and barley pathogens in airborne samples
- Detection preceded visible symptom appearance in experimental conditions
- The method simultaneously detected plant pathogens, insect pests, and background biodiversity from a single analysis
- Comparison with traditional Burkard trap monitoring showed similar or better performance for fungal detections
2.2 Giolai et al. (2024): air metagenomics in an agricultural ecosystem
Giolai et al. (2024) characterised the air metagenome of an agricultural ecosystem using Oxford Nanopore long-read sequencing. They identified diverse plant pathogens, beneficial microorganisms, and crop species DNA in the air of active agricultural fields. Long-read sequencing provided better resolution of fungal taxa compared to short-read metabarcoding, which is important for distinguishing closely related pathogen strains.
2.3 Comparison with traditional methods (Berelson et al. 2025)
Berelson et al. (2025) reviewed comparisons between traditional spore trap microscopy and molecular metabarcoding on the same samples. Molecular methods detected pathogenic species that microscopy missed, because some pathogen spores have overlapping morphology with non-pathogenic congeners that only DNA sequencing can distinguish. This provides a direct accuracy argument for the molecular approach over the traditional one.
3. The agricultural infrastructure opportunity
Agriculture already has airborne monitoring infrastructure that is potentially deployable for eDNA surveillance:
Burkard volumetric spore traps: Already deployed at many sites across the UK and Europe to monitor pollen and pathogen spores for allergy forecasting and crop disease advisory services. These devices continuously sample air onto adhesive tape and produce archived biological material that is directly amenable to DNA extraction and sequencing.
AHDB Crop Monitor (UK): The Agriculture and Horticulture Development Board already operates a crop disease monitoring network that combines field scouting, spore trap data, and meteorological modelling to produce disease risk forecasts. Adding molecular pathogen detection to this existing system would represent an incremental step rather than a new programme.
For the Netherlands and broader Europe, equivalent infrastructure exists. Naktuinbouw (the Dutch Plant Protection Service) and Wageningen University & Research both operate crop disease monitoring and advisory services, and the European and Mediterranean Plant Protection Organisation (EPPO) coordinates pan-European pest and pathogen surveillance through its reporting service. Several Burkard volumetric trap networks are embedded within European pollen and spore monitoring frameworks (including those coordinated by the European Aerobiology Society), which have already been used for crop pathogen tracking. The integration of molecular eDNA analysis into these existing European networks represents a similarly incremental opportunity.
Meteorological warning integration: Pathogen spore dispersal is usually weather-dependent (following rain events and during periods of wind from known source areas). Integrating airborne eDNA detection with HYSPLIT back-trajectory analysis could identify not just that a pathogen is present but where it likely came from — enabling source-targeted management responses.
4. The insect pest application
Beyond fungal pathogens, the detection of insect agricultural pests via airborne eDNA has significant potential. Roger et al. (2022) and Pumkaeo et al. (2021) demonstrated detection of insect taxa from airborne eDNA. For invasive pest monitoring — detecting the first arrival of a new pest species in a region — the non-targeted surveillance capability of airborne eDNA metabarcoding offers a genuine advance over species-specific sentinel trapping.
The EU Regulation on Protective Measures against Pests of Plants (Plant Health Regulation 2016/2031) requires member states to maintain surveillance for priority pests. Airborne eDNA could provide a cost-effective, scalable surveillance layer for the list of regulated EU quarantine pests, operating continuously at sites where conventional field-based surveillance is logistically difficult or financially prohibitive.
5. Nanopore sequencing for field-deployable, real-time detection
One of the most exciting developments for the agricultural application is the potential for field-deployable, real-time molecular diagnosis. Oxford Nanopore’s MinION device (Jain et al. 2016) is roughly the size of a mobile phone and produces sequencing results within hours of sample preparation. Combined with a portable air sampler, this creates the possibility of a self-contained field unit that a well trained agricultural advisor, or agronomist could deploy in a field to detect emerging pathogen threats in near real-time.
Berelson et al. (2025) identify this as a key emerging application, noting that real-time pathogen detection has transformative potential for precision agriculture and food security.
6. Barriers to operational deployment
Sensitivity thresholds for early warning. How low a spore concentration can the method reliably detect? At what point before symptom expression can detection be made? These parameters need to be characterised for each pathogen species of concern before the system can be trusted for decision support.
Reference database coverage for agricultural pathogen strains. Many important crop pathogens have limited reference sequence coverage, particularly at the strain level. Different strains may have different fungicide resistance profiles; strain-level detection is needed for optimising treatment selection.
Integration with agronomic decision frameworks. A detection alone does not constitute a treatment recommendation. Integration with disease risk models, field history records, fungicide resistance management guidelines, and economic thresholds is needed before airborne eDNA data can feed directly into crop management decisions.
Regulatory pathway for data use. In the UK, crop disease advisory services operate within a regulatory and liability framework. The use of airborne eDNA data in management decisions needs an agreed pathway for data validation, quality assurance, and liability allocation.
References
- FAO. Plant production and protection. Food and Agriculture Organization of the United Nations. https://www.fao.org/plant-production-protection/about/en
- Mikko A et al. (2025). Sequencing airborne DNA to monitor crop pathogens and pests. iScience. https://doi.org/10.1016/j.isci.2025.112912
- Nousias O et al. (2025). Shotgun sequencing of airborne eDNA achieves rapid assessment of whole biomes. Nature Ecology & Evolution. https://doi.org/10.1038/s41559-025-02711-w
- Giolai M et al. (2024). Measuring air metagenomic diversity in an agricultural ecosystem. Current Biology. https://doi.org/10.1016/j.cub.2024.07.030
- Berelson MFG et al. (2025). From air to insight: the evolution of airborne DNA sequencing technologies. Microbiology 171:001564. https://doi.org/10.1099/mic.0.001564
- Roger F et al. (2022). Airborne environmental DNA metabarcoding for the monitoring of terrestrial insects. Environmental DNA 4:790–807. https://doi.org/10.1002/edn3.290
- Jain M et al. (2016). The Oxford Nanopore MinION: delivery of nanopore sequencing to the genomics community. Genome Biology 17:239. https://doi.org/10.1186/s13059-016-1103-0
- Pumkaeo P et al. (2021). Detection and monitoring of insect traces in bioaerosols. PeerJ 9:e10862. https://doi.org/10.7717/peerj.10862