Short Answer: We do not yet have a precise answer to how long airborne eDNA persists. Under harsh outdoor conditions — high UV, summer heat, exposed particle surfaces — DNA can degrade to below-detection within hours. Under protective conditions — DNA enclosed within an intact pollen grain or spore, cool temperatures, overcast conditions — it can survive for days to weeks. The practical reality for most field vertebrate monitoring is that a detection most likely reflects presence within the past few hours to a few days. Critically, this uncertainty is not yet incorporated into the atmospheric transport models used to trace signal origins — meaning modelled catchment areas represent where the particles could physically have come from, not necessarily where detectable DNA actually originated.
1. Why persistence matters
The question of DNA persistence in air determines a critical interpretation: when a species is detected in an air sample, how recently was it present? This matters for:
2. The mechanisms of DNA degradation in air
DNA in the atmosphere is subject to several degradation pathways simultaneously:

2.1 UV photodegradation
Ultraviolet radiation — particularly UV-B (280–315 nm) — is the most potent driver of outdoor DNA degradation. UV photons induce the formation of pyrimidine dimers in DNA strands, which block replication and transcription enzymes. At sufficient doses, UV radiation fragments DNA into pieces too small to amplify by PCR.
Biological particles vary in their UV resistance depending on whether their DNA is encapsulated or exposed. Intact pollen grains and bacterial cells have physical and chemical shields (sporopollenin in pollen; protective protein coats in bacteria) that substantially reduce UV penetration. Naked DNA fragments from cell debris are far more vulnerable. Backes et al. (2021) documented that chemical oxidation and nitration by atmospheric oxidants (ozone, nitrogen dioxide, peroxynitrite) modify pollen allergen proteins — work focused on allergenicity rather than eDNA degradation, but it illustrates the chemically reactive environment airborne biological particles inhabit.
2.2 Temperature effects
High temperatures accelerate all chemical degradation reactions. In summer conditions with temperatures exceeding 30°C and full UV exposure, DNA in unprotected particles may degrade within hours. In winter conditions with temperatures near 0°C and reduced UV, degradation is much slower.
2.3 Enzymatic degradation
Extracellular DNA (DNA outside intact cells) is rapidly degraded by DNase enzymes produced by bacteria, which are themselves present in the atmosphere. The rate of enzymatic degradation depends on atmospheric humidity, temperature, and the density of microbial communities on particle surfaces. In hot, humid environments, enzymatic degradation can be rapid. In cool, dry conditions or inside protective cellular structures, it is greatly slowed.
2.4 Physical fragmentation
Mechanical stresses during atmospheric transport — turbulence, particle-particle collisions, impaction on surfaces — can physically fragment DNA molecules. This is particularly relevant for very long DNA molecules; shorter fragments (as targeted by most eDNA metabarcoding primers) are more resistant to physical fragmentation.
3. What the evidence says about persistence timescales
3.1 Short-term persistence in field conditions
Direct measurement of airborne DNA persistence under field conditions is surprisingly rare. Most inference is indirect — drawn from temporal patterns of detection rather than controlled degradation experiments.
The weather conditions that drive particle transport also shape degradation rate, but through different mechanisms. Wind moves particles; UV radiation and heat destroy their DNA content. This creates a fundamental asymmetry: a highly transported particle from far away may arrive with degraded DNA, while a freshly shed particle nearby may be detectable for only a short window before UV exposure renders it undetectable. Weather data collected alongside air samples helps interpret these dynamics, but cannot yet disentangle transport time from degradation rate for any individual detection.
For aquatic eDNA — the best-studied system — DNA in flowing water typically persists for hours to a few days before degradation or dilution below detection thresholds (Jane et al. 2015; Harrison et al. 2019). Airborne DNA exposed to outdoor UV is likely to degrade at least as fast, and potentially faster, than water eDNA under comparable conditions.
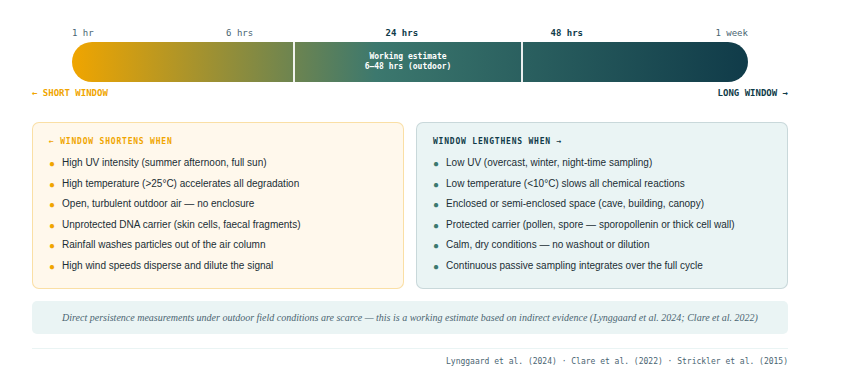
A working estimate for outdoor field conditions under moderate UV exposure: airborne vertebrate eDNA likely represents the past 6–48 hours, with high variability depending on temperature, UV intensity, and whether the DNA is encapsulated in intact cells.
3.2 Persistence in enclosed and open environments
The most informative persistence data come from enclosed and semi-enclosed environments, where transport is constrained and persistence can be more directly studied. Clare et al. (2021) demonstrated that DNA accumulated in sealed indoor air and remained detectable hours after source animals had been removed. Clare et al. (2022) confirmed this from zoo enclosures: DNA persisted inside sealed buildings at low-UV, low-turbulence conditions for at least several hours.
Garrett et al. (2025) pushed this further with controlled experiments in a semi-enclosed field setting — a room-sized space with known faunal composition — finding that detection probability varied with both sampling duration and the number of samplers deployed. Species present but rarely detected in short samples became consistently detectable with longer deployment, suggesting that the temporal integration window matters substantially for low-abundance targets. In enclosed spaces where DNA accumulates rather than dispersing, sampling duration is a direct lever on detection completeness.
In open natural landscapes the same principles apply but with two important counteracting forces: atmospheric mixing dilutes DNA concentrations faster, and UV exposure is greater, shortening the persistence window. A detection in an enclosed building is more likely to reflect an animal present within the past few hours; a detection in an open field integrates a somewhat shorter and harder-to-quantify window. For rare single-sample observations from open landscapes — particularly those where long-range transport is possible — the temporal origin of the detected DNA cannot be specified with precision, and this uncertainty should be acknowledged when interpreting the result.
4. How particle type shapes persistence
Not all airborne DNA persists equally. The type of particle carrying the DNA dramatically affects its persistence:
| Carrier particle | UV protection | Typical indoor persistence | Typical outdoor persistence |
|---|---|---|---|
| Intact pollen grain | High (sporopollenin wall) | Days to weeks | Hours to days |
| Intact bacterial cell | Moderate (cell wall, pigments) | Days | Hours |
| Intact fungal spore | High (thick cell wall) | Days to weeks | Hours to days |
| Fragmented cell debris | Low (no structural protection) | Hours | Minutes to hours |
| Dried faecal particle | Moderate (organic matrix) | Days | Hours |
Pollen DNA is particularly persistent because sporopollenin — the material forming pollen walls — is one of the most chemically resistant biological polymers known. Pollen extracted from ancient sediments retains identifiable morphology for millions of years. By contrast, naked DNA fragments from shed skin cells or faecal aerosols are exposed to the environment and degrade much faster.
This particle-type variation means that plant and fungal eDNA likely persists longer in air samples than vertebrate or invertebrate eDNA. A monitoring programme focused on vertebrate detection may be capturing a shorter temporal window than one focused on plant or fungal communities.
5. Caveats and open questions
Direct persistence experiments under outdoor conditions are lacking. Unlike aquatic eDNA (which has been the subject of controlled degradation experiments under defined UV, temperature, and flow conditions: Strickler et al. 2015; Bylemans et al. 2018), outdoor airborne eDNA persistence has not been measured in controlled field experiments. This is a significant gap that limits quantitative interpretation of detection timing.
Persistence varies by taxon, particle type, and environment in ways not yet predictable. The current understanding does not allow practitioners to specify, for a given detection, whether the DNA is 2 hours or 24 hours old.
Atmospheric transport models do not yet account for DNA degradation. Models such as HYSPLIT and FLEXPART simulate the physical movement of particles from potential source regions to receptor locations. They can estimate the geographic area from which particles could physically have arrived. But they do not incorporate DNA decay rates — meaning they cannot currently distinguish between particles that arrived with intact, detectable DNA and particles that arrived degraded beyond detection. The modelled catchment therefore represents a physical source envelope, not a detectable source envelope. In practice the detectable catchment is smaller — constrained by how long the DNA survived in transit. Quantifying this gap requires integrating degradation kinetics into dispersion models, which has not yet been done for macrobial airborne eDNA. It is a critical next step for converting model outputs into biologically interpretable source attributions.
References
- Backes AT et al. (2021). Oligomerization and nitration of the grass pollen allergen Phl p 5 by ozone, nitrogen dioxide, and peroxynitrite. International Journal of Molecular Sciences 22:7616. https://doi.org/10.3390/ijms22147616
- Bálint M et al. (2018). Environmental DNA time series in ecology. Trends in Ecology & Evolution 33:945–957. https://doi.org/10.1016/j.tree.2018.09.003
- Bylemans J et al. (2018). Does size matter? An experimental evaluation of the relative abundance and decay rates of aquatic environmental DNA. Environmental Science & Technology 52:6408–6416. https://doi.org/10.1021/acs.est.8b01071
- Clare EL et al. (2021). eDNAir: proof of concept that animal DNA can be collected from air sampling. PeerJ 9:e11030. https://doi.org/10.7717/peerj.11030
- Clare EL et al. (2022). Measuring biodiversity from DNA in the air. Current Biology 32:693–700. https://doi.org/10.1016/j.cub.2021.11.064
- Garrett NR et al. (2025). Sampling intensity and temporal persistence of airborne eDNA in partially enclosed spaces. bioRxiv preprint. https://doi.org/10.1101/2025.07.14.664745
- Harrison JB et al. (2019). Predicting the fate of eDNA in the environment. Proceedings of the Royal Society B 286:20191409. https://doi.org/10.1098/rspb.2019.1409
- Jane SF et al. (2015). Distance, flow, and PCR inhibition: eDNA dynamics in two headwater streams. Freshwater Biology 60:2382–2394. https://doi.org/10.1111/fwb.12523
- Lynggaard C et al. (2024). Airborne environmental DNA captures terrestrial vertebrate diversity in nature. Molecular Ecology Resources 24:e13840. https://doi.org/10.1111/1755-0998.13840
- Strickler KM et al. (2015). Quantifying effects of UV-B, temperature, and pH on eDNA degradation. Biological Conservation 183:85–92. https://doi.org/10.1016/j.biocon.2014.11.038