Short Answer: Contamination is almost an unavoidable risk in airborne eDNA work, but it is manageable with protocols. The main sources are: DNA from laboratory workers or environment entering samples (human contamination); carryover between samples in the lab (cross-contamination); DNA from the sampling environment entering samples during collection (field contamination); and errors in the reference database (informational contamination). Each has established mitigation strategies. A credible airborne eDNA study must explicitly document and control for contamination at every stage.
1. Why contamination is a special concern in airborne eDNA
All eDNA methods are highly sensitive to contamination because the analytical method — PCR — amplifies tiny amounts of DNA by factors of millions. A single stray molecule can, in theory, produce a positive signal. The concentration of target DNA in many airborne samples (particularly from rare or distant taxa) is very low, making the ratio of signal to potential contamination smaller than in water eDNA from a productive river or lake.
Additionally, the very principle of airborne eDNA sampling — collecting DNA from the air — means that the sampling environment is the atmosphere, which contains biological particles from many sources including the humans doing the sampling. This creates a ubiquitous and ambient contamination risk that affects soil or water sampling to a lesser extent.
2. Field contamination: DNA from the sampling environment
2.1 Sampler contamination from handlers
The person deploying, retrieving, or handling an air sampler inevitably sheds DNA (skin cells, breath aerosols, hair) in the vicinity of the sampler. If this DNA is deposited on the filter during handling, it will appear as a human detection and could also physically displace or mask non-human DNA.
Mitigation: Handle all sampler components wearing nitrile gloves and, where possible, a face mask and hair cover. Minimise the time filters are exposed to ambient air during handling. Prepare filters in a clean laboratory environment before deployment.
More critically, human handlers can introduce non-local eDNA from any location they have recently visited. A field technician who has been on a different survey site that morning — whether a nature reserve, a farm, or a wetland — may carry viable DNA from those environments on their clothing, equipment, or skin. This is distinct from simply shedding human DNA: it means that species genuinely absent from the survey area can appear as detections if handling protocols are not rigorous. For this reason, aquatic eDNA protocols developed by agencies such as the US Fish and Wildlife Service require technicians to decontaminate waders, boots, and sampling equipment between sites using bleach or isopropanol solutions, and to record their recent site history so any cross-contamination can be traced. Equivalent standards may have to be applied in airborne eDNA work, but practically one may also have to accept a level of contamination risk when interpreting results and planning campaigns.
2.2 Ambient lab contamination during filter preparation
Filters prepared in a laboratory where other biological samples are processed risk contamination from DNA already present in the lab environment. Clare et al. (2022) prepared filters in dedicated eDNA labs with UV sterilisation between sessions and strict decontamination of all equipment.
2.3 Transport contamination
Filters transported to and from the field without sealed packaging can accumulate environmental DNA from transit environments. For this reason a survey often deploys field blanks. These samples are handled through the full deployment and collection process but not actually exposed to the target air. Any species detected in a field blank must be treated as a potential false positive for the associated samples.
3. Laboratory contamination: the dominant risk
3.1 Cross-contamination between samples
DNA from high-concentration samples can contaminate adjacent low-concentration samples during processing. This is particularly problematic when processing multiple filters from the same monitoring network: a filter with high signal from a common species can contaminate a filter that should have shown no signal for that species.
Mitigation: Process samples in small batches; include blank filter controls between samples; use physical separation between positive samples and controls; UV-treat work surfaces between samples.
3.2 PCR carryover
PCR amplification produces enormous quantities of amplified DNA (amplicons). If amplicons from one reaction enter the reaction mixture of another sample — through aerosols, contaminated pipettes, or open tube handling — they produce false positive signals. This is called “carryover contamination” and is the most serious laboratory contamination risk.
Mitigation: Physically separate pre- and post-PCR areas; use dedicated pipettes for each area; use dUTP/UNG (uracil-N-glycosylase) enzymatic decontamination where possible; include PCR negative controls in every run.
3.3 Laboratory environment contamination
Molecular biology laboratories accumulate background DNA from years of previous experiments. Regular decontamination with bleach (sodium hypochlorite), UV irradiation, and commercial DNA decontaminants (e.g. DNAaway) is essential.
Clare et al. (2022) conducted all DNA extraction and PCR in a biological safety cabinet under maximum flow, sterilised equipment with UV, 10% bleach, 70% ethanol and ultrapure water between each sample, and used multiple negative controls at extraction, PCR and sequencing stages.
4. Human DNA contamination: a special category
Human DNA is a particularly important contaminant in airborne eDNA work because:
-
1Humans are always present at sampling sites and in laboratories
-
2Human DNA primers are present in some universal mammalian primer panels
-
3Detected human DNA can create legal, ethical and privacy concerns
Nousias et al. (2025) and the Science AAAS coverage of airborne eDNA both highlighted that shotgun sequencing of air samples captures human genomic DNA from people present in or near the sampling location. This is not contamination in the analytical sense — it is a genuine detection of human eDNA. But it raises questions about consent, privacy, and the appropriate handling of human genetic data incidentally collected during environmental monitoring.
Mitigation for analytical purposes: Use primers or bioinformatic filters that exclude human sequences from analysis. For metabarcoding, this is straightforward (human sequences are simply not assigned to environmental taxa). For shotgun metagenomics, human reads should be filtered against a human reference genome before ecological analysis.
5. Informational contamination: database errors
A form of contamination that is less discussed but equally important is errors in reference databases. If a species is incorrectly identified in the reference database, or if a sequence from one species is accidentally deposited under another species’ name, all eDNA detections matching that erroneous entry will be misidentified. This is not a contamination of the physical sample, but of the informational chain that converts sequence to species name.
Mitigation: Use curated, high-quality reference databases rather than unfiltered general repositories; cross-check unexpected detections against multiple databases; build local reference collections for focal taxa.
6. Control samples: the backbone of quality assurance
It is recommended that airborne eDNA studies include a systematic set of negative and positive controls.
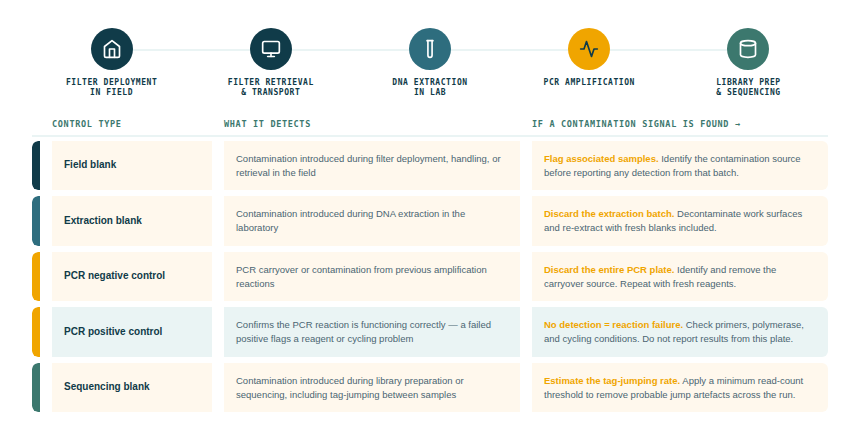
Any species detection appearing in a negative control must be investigated and, if not explicable, must cause the associated samples to be flagged or excluded. The level of contamination acceptable depends on the application.
7. Decontamination protocols
A minimum decontamination protocol for airborne eDNA work includes:
- Filter handlingGloves, mask, hair cover; forceps decontaminated between samples; filters stored individually in sealed bags
- Work surfacesUV irradiation for 30+ minutes; wipe down with 10% bleach followed by 70% ethanol and ultrapure water
- EquipmentDedicated sets for pre- and post-PCR work; single-use consumables where possible
- PipettesFilter tips only; calibrated and decontaminated regularly
- PersonnelGloves changed frequently; no food, drink or bare-skin contact in the eDNA processing area
Clare et al. (2022) described a decontamination and extraction protocol for vertebrate airborne eDNA — using a biological safety cabinet under maximum flow, UV sterilisation, 10% bleach, 70% ethanol and ultrapure water between each sample, and multiple negative controls at extraction, PCR and sequencing stages — that has been referenced and adapted by a number of subsequent research groups as a starting point for their own protocols. It should be noted that the field has not yet converged on a single standard: different laboratories use variations of this approach, and comparisons between studies are complicated by these methodological differences.
8. Implications for regulatory use
For regulatory applications — species protection surveys, planning assessments, national monitoring programmes — contamination control protocols must be:
The great crested newt eDNA protocol in the UK (Biggs et al. 2015) provides a template for this level of protocol documentation. An equivalent framework for airborne eDNA regulatory applications does not yet exist but is needed as the method moves towards routine deployment.
References
- Alberdi A et al. (2018). Scrutinizing key steps for reliable metabarcoding. Methods in Ecology and Evolution 9:134–147. https://doi.org/10.1111/2041-210x.12849
- Berelson MFG et al. (2025). From air to insight. Microbiology 171:001564. https://doi.org/10.1099/mic.0.001564
- Biggs J et al. (2015). Using eDNA to develop a national citizen science-based monitoring programme for the great crested newt (Triturus cristatus). Biological Conservation 183:19–28. https://doi.org/10.1016/j.biocon.2014.11.029
- Clare EL et al. (2022). Measuring biodiversity from DNA in the air. Current Biology 32:693–700. https://doi.org/10.1016/j.cub.2021.11.064
- Goldberg CS et al. (2016). Critical considerations for the application of environmental DNA methods to detect aquatic species. Methods in Ecology and Evolution 7:1299–1307. https://doi.org/10.1111/2041-210X.12595
- Nousias O et al. (2025). Shotgun sequencing of airborne eDNA achieves rapid assessment of whole biomes. Nature Ecology & Evolution. https://doi.org/10.1038/s41559-025-02711-w
- Tournayre O et al. (2025). First national survey of terrestrial biodiversity using airborne eDNA. Scientific Reports. https://doi.org/10.1038/s41598-025-03650-z