Short Answer: Every living organism — from bacteria to elephants — continuously sheds fragments of DNA into its surroundings. In terrestrial environments, much of this genetic material becomes airborne, attaching to microscopic particles that float in the atmosphere. By filtering air samples and sequencing the DNA they contain, scientists can identify which species are present in an area without ever seeing, trapping or handling a single animal or plant. This is airborne environmental DNA.
1. The concept: DNA as an ecological signal
Every organism on Earth is a continuous source of genetic material. Cells are shed through skin, fur, feathers, faeces, urine, saliva, pollen, spores, and the simple process of decomposition. In aquatic ecosystems, this shed DNA — termed environmental DNA, or eDNA — dissolves or suspends in the water column, where it can be captured by filtering a water sample. The power of this approach transformed freshwater ecology over the past two decades: a single litre of river water can yield genetic evidence of dozens of species without a net, trap or camera ever being deployed (Thomsen & Willerslev 2015; Deiner et al. 2017).
The conceptual leap that underpins airborne eDNA is straightforward: air, like water, is a medium through which biological particles travel. The near-surface atmosphere is not empty — it is a complex suspension of organic particles including fungal spores, pollen grains, bacterial cells, plant fragments, animal hair, skin cells, and faecal particles. Embedded within all of these are DNA molecules. If those molecules can be captured and sequenced, the air itself becomes a biodiversity sensor.
This idea was formulated independently and demonstrated simultaneously by two research groups in early 2022, published on the same day in Current Biology (Clare et al. 2022; Lynggaard et al. 2022). Both groups collected air samples at European zoos — where the species composition was known — and detected DNA from dozens of mammals and birds. The independent replication of the same finding by teams with no knowledge of each other's work substantially strengthened the credibility of the approach. As Kristine Bohmann, lead of the Danish study, remarked: "We did not think that vacuuming animal DNA from air would work. This was high risk, high reward science."
2. What organisms shed DNA into the air?
The biological sources of airborne eDNA span the entire tree of life. Five broad groups are detectable, each shedding DNA through distinct mechanisms. This section gives an introduction to each; the following sections cover what has been detected, how reliably, and what the practical implications are by taxon group.

Vertebrates — mammals and birds shed DNA passively through skin cells, hair, feathers, breath aerosols, and faeces. Unlike plants (which release pollen) or fungi (which release spores), vertebrates have no dedicated airborne dispersal stage: they simply shed detectable quantities of DNA by existing in an environment, breathing and moving. The earliest zoo studies (Clare et al. 2022; Lynggaard et al. 2022) detected species through sealed buildings at hundreds of metres, establishing that vertebrate signal is robust.
Plants — wind-pollinated species shed pollen, the focus of aerobiology for decades. More recently, even insect-pollinated plants that produce little airborne pollen have been detected via leaf cuticle fragments and other non-pollen particles released by wind action, particularly during storms (Johnson et al. 2019; Hanson et al. 2024). Plant community monitoring via air is not restricted to the pollen season.
Fungi — spores are among the most abundant biological particles in near-surface air, routinely exceeding 10,000 per cubic metre during peak seasons, and present year-round. Fungal eDNA has the longest monitoring history of any airborne taxon group, with an established global dataset from the Global Spore Sampling Project (Ovaskainen et al. 2024).
Insects and invertebrates — detectable via shed scales, cuticle fragments, frass, and body debris. The first rigorous proof of concept came from Roger et al. (2022); multiple field studies have since confirmed detection across arthropod communities. Given the documented collapse of flying insect populations in many regions (Hallmann et al. 2017), scalable monitoring via air filters is of considerable conservation significance.
Bacteria and microorganisms — microbial eDNA in air has been studied for decades under aerobiology and atmospheric microbiology. Bacterial concentrations typically range from 10² to 10⁴ cells per cubic metre. Sullivan et al. (2025) detected over 2,700 genera spanning prokaryotes and eukaryotes from 34 years of archived aerosol filters, demonstrating the atmospheric archive extends across all domains of life.
The physical form these particles take — and why it matters for both detection and interpretation — is covered in section 3 below.
3. The physical form of airborne eDNA
Understanding what airborne eDNA is physically — not just biologically — is essential for interpreting detections and designing sampling strategies. This is where insights from aerosol physics and atmospheric science become critical, and where the airborne eDNA field has much to learn from neighbouring disciplines.
3.1 Particle size classes
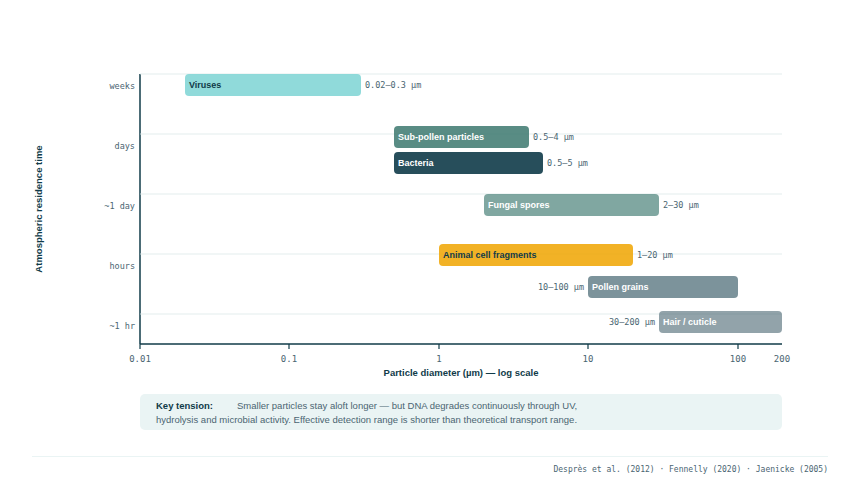
Biological particles in the atmosphere span an enormous size range:
| Particle type | Typical diameter | Atmospheric behaviour |
|---|---|---|
| Viruses | 0.02–0.3 µm | Long-range transport possible |
| Bacteria | 0.5–5 µm | Days to weeks aloft |
| Fungal spores | 2–30 µm | Hours to days aloft |
| Sub-pollen particles | 0.5–4 µm | Days aloft |
| Pollen grains | 10–100 µm | Hours to days; gravity limits range |
| Animal cell fragments | 1–20 µm | Hours aloft |
Smaller particles remain airborne longer and travel further. The largest pollen grains (e.g. pine, 50–100 µm) deposit within hours of release, while sub-pollen particles and bacterial cells can remain aloft for days and travel thousands of kilometres (Desprès et al. 2012; Jaenicke 2005; Prenni et al. 2009). This has direct implications for interpreting airborne eDNA detections: the particle size of the DNA carrier partly determines the geographic catchment area of a sample. However, this must be weighed against the counteracting effect of DNA degradation: small particles may travel far, but the DNA they carry degrades continuously through UV radiation, hydrolysis, and microbial activity. Atmospheric residence time and DNA integrity are therefore in tension — a particle capable of travelling days may carry no detectable sequence by the time it arrives. The effective detection range is substantially shorter than the theoretical transport range.
3.2 DNA on particles vs free DNA
DNA in air is not expected to be present as 'naked', free-floating molecules — it is almost always associated with a carrier particle. The carrier may be the intact biological cell itself or fungal spore, with the DNA still enclosed within, whilst free DNA is likely to adhere to solid particles or be encapsulated in aerosols. This is significant for two reasons. First, association with a solid particle provides physical protection from UV radiation, which is one of the primary drivers of DNA degradation. Second, the type of particle determines how the DNA behaves atmospherically — its deposition velocity, response to humidity, and susceptibility to wet scavenging by rain.
Fröhlich-Nowoisky et al. (2016) established the concept of primary biological aerosol particles (PBAPs) as a distinct category within atmospheric aerosol science — particles that are directly emitted from biological sources, as opposed to secondary biological aerosols formed by chemical reactions in the atmosphere. This distinction matters for eDNA practitioners: the DNA they detect is associated with PBAPs, not with inorganic dust or chemical aerosols.
4. Why air, and why now?
4.1 The global biodiversity monitoring gap
The scale of the biodiversity crisis is well documented but poorly quantified. Conventional monitoring methods — transect surveys, camera trapping, acoustic monitoring, light trapping for insects — are labour-intensive, taxonomically specialised, geographically patchy, and temporally discontinuous. Recently acoustic monitoring is advancing rapidly though and does offer valuable continuous temporal resolution for some taxa; its evolving capabilities are a complement to the landscape-scale taxonomic breadth that eDNA can provide. The WWF Living Planet Report estimates a 69% average decline in vertebrate populations since 1970, but these estimates are built from datasets that cover a tiny and non-random fraction of global biodiversity (Ceballos et al. 2020; Newbold et al. 2015).
Airborne eDNA addresses this gap in a specific and important way: it is taxonomically non-selective, geographically scalable, and compatible with existing infrastructure. A single air filter processed with shotgun sequencing can yield information about vertebrates, plants, fungi, insects and microorganisms simultaneously — without prior knowledge of which taxa are present and without taxonomic expertise in the field (Sullivan et al. 2025; Tournayre et al. 2025).
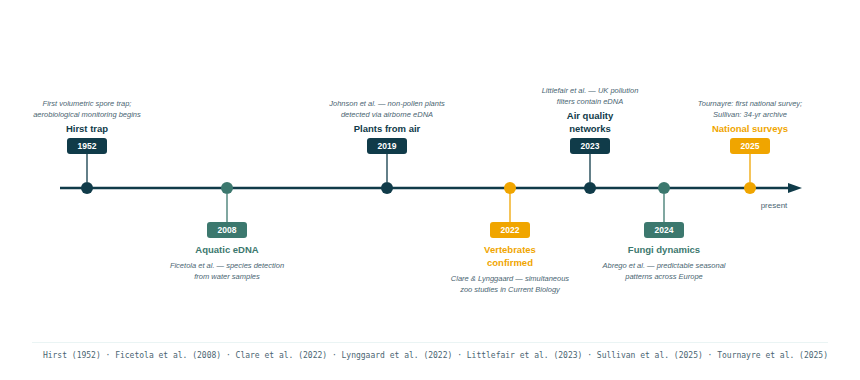
4.2 A field coming of age
Publication rates in airborne macrobial eDNA grew from near zero before 2019 to dozens of papers per year by 2024–2025 (Johnson & Barnes 2024). The field is no longer at proof-of-concept stage for vertebrates and plants. What remains to be standardised — and what prevents regulatory adoption — are the methodological protocols, quality controls, and interpretive frameworks needed to make airborne eDNA data legally defensible and scientifically comparable across sites and time (Tulloch et al. 2025; NPL/LGC 2025).
5. Key caveats
Two caveats are important enough to flag at the outset of any engagement with airborne eDNA:
DNA does not equal organism. Detecting a species' DNA in the air does not confirm that the species is currently present at the sampling location. DNA can travel long distances, persist for hours to days, and be shed by animals passing through rather than residing. The nuances of what a detection does and does not imply are addressed in detail in the final section of this article on caveats.
Method standardisation is still developing. Different sampler types, filter materials, extraction protocols and sequencing strategies yield different results from the same air. Without agreed reference methods, datasets from different studies cannot be reliably compared. This is the field's most urgent practical challenge (Tulloch et al. 2025), and affects what airborne eDNA can and cannot currently be used for in formal regulatory contexts.
References
- Abrego N et al. (2024). Airborne DNA reveals predictable spatial and seasonal dynamics of fungi. Nature 631(8022):835–842. https://doi.org/10.1038/s41586-024-07658-9
- Clare EL et al. (2022). Measuring biodiversity from DNA in the air. Current Biology 32(3):693–700. https://doi.org/10.1016/j.cub.2021.11.064
- Deiner K et al. (2017). Environmental DNA metabarcoding: Transforming how we survey animal and plant communities. Molecular Ecology 26:5872–5895. https://doi.org/10.1111/mec.14350
- Desprès VR et al. (2012). Primary biological aerosol particles in the atmosphere: A review. Tellus B 64:15598. https://doi.org/10.3402/tellusb.v64i0.15598
- Fennelly KP (2020). Particle sizes of infectious aerosols: implications for infection control. The Lancet Respiratory Medicine 8(9):914–924. https://doi.org/10.1016/S2213-2600(20)30323-4
- Ficetola GF et al. (2008). Species detection using environmental DNA from water samples. Biology Letters 4:423–425. https://doi.org/10.1098/rsbl.2008.0118
- Fröhlich-Nowoisky J et al. (2016). Bioaerosols in the Earth system: Climate, health, and ecosystem interactions. Atmospheric Research 182:346–376. https://doi.org/10.1016/j.atmosres.2016.07.018
- Hallmann CA et al. (2017). More than 75 percent decline over 27 years in total flying insect biomass. PLOS ONE 12:e0185809. https://doi.org/10.1371/journal.pone.0185809
- Hanson M et al. (2024). Storms facilitate airborne DNA from leaf fragments outside the main tree pollen season. Aerobiologia 40(3):415–423. https://doi.org/10.1007/s10453-024-09826-w
- Hirst JM (1952). An automatic volumetric spore trap. Annals of Applied Biology 39:257–265. https://doi.org/10.1111/j.1744-7348.1952.tb00904.x
- Jaenicke R (2005). Abundance of cellular material and proteins in the atmosphere. Science 308:73. https://doi.org/10.1126/science.1106335
- Johnson M & Barnes MA (2024). Macrobial airborne environmental DNA analysis: A review of progress, challenges, and recommendations. Molecular Ecology Resources 24:e13998. https://doi.org/10.1111/1755-0998.13998
- Littlefair JE et al. (2023). Air-quality networks collect environmental DNA with the potential to measure biodiversity at continental scales. Current Biology 33:R426–R428. https://doi.org/10.1016/j.cub.2023.04.036
- Lynggaard C et al. (2022). Airborne environmental DNA for terrestrial vertebrate community monitoring. Current Biology 32(3):701–707. https://doi.org/10.1016/j.cub.2021.12.014
- NPL/LGC (2025). Standardising airborne eDNA for biodiversity monitoring: a UK collaboration to advance measurement science. National Physical Laboratory / National Measurement Laboratory at LGC. https://www.uknml.com/resources/standardising-airborne-edna-for-biodiversity-monitoring-a-uk-collaboration-to-advance-measurement-science-2/
- Roger F et al. (2022). Airborne environmental DNA metabarcoding for the monitoring of terrestrial insects. Environmental DNA 4:790–807. https://doi.org/10.1002/edn3.290
- Sofiev M et al. (2013). A numerical model of birch pollen emission and dispersion in the atmosphere. International Journal of Biometeorology 57:45–58. https://doi.org/10.1007/s00484-012-0532-z
- Sullivan AR et al. (2025). Airborne eDNA captures three decades of ecosystem biodiversity. Nature Communications 16:11281. https://doi.org/10.1038/s41467-025-67676-7
- Thomsen PF & Willerslev E (2015). Environmental DNA – An emerging tool in conservation for monitoring past and present biodiversity. Biological Conservation 183:4–18. https://doi.org/10.1016/j.biocon.2014.11.019
- Tournayre O et al. (2025). First national survey of terrestrial biodiversity using airborne eDNA. Scientific Reports 15:19247. https://doi.org/10.1038/s41598-025-03650-z
- Tulloch RL et al. (2025). Winds of Change: Charting a Pathway to Ecosystem Monitoring Using Airborne Environmental DNA. Environmental DNA 7:e70134. https://doi.org/10.1002/edn3.70134