Short Answer: Although no standard method exists, airborne eDNA is usually collected by drawing air through a filter or by allowing DNA-laden particles to settle passively onto a collecting surface. The filter is then processed in a laboratory to extract and amplify the DNA, which is sequenced and matched against reference databases to identify species. The full pipeline from field to species list now takes days to weeks and costs considerably less than traditional biodiversity surveys over comparable areas, though no single standardised protocol yet exists.
1. The basic pipeline
Every airborne eDNA study, regardless of the specific equipment used, follows the same general workflow:
-
1Air samplingbiological particles are concentrated and captured
-
2DNA extractionDNA is chemically liberated from the collected material
-
3Amplificationtarget DNA sequences are copied using PCR (polymerase chain reaction)
-
4Sequencingamplified DNA is sequenced using next-generation or nanopore technology
-
5Bioinformatic analysissequences are matched against reference databases to assign taxonomy
-
6Ecological interpretationdetections are translated into species presence/absence or relative abundance
The quality of the final output depends critically on decisions made at every step. Errors or biases introduced during sampling propagate through the entire pipeline and cannot be corrected computationally downstream (Berelson et al. 2025; Alberdi et al. 2018).
2. Air sampling methods: active vs passive
The most visible methodological choice in airborne eDNA study design is between active and passive sampling — but this is only one of several critical decisions. Sampling design also encompasses filter type and pore size, deployment duration, spatial replication, and whether the primary aim is targeted detection of a known species (favouring qPCR-based approaches) or community-wide surveillance. The active/passive distinction remains a useful organising principle, but practitioners should treat it as one axis in a multi-dimensional design space, not as the sole determining factor.
⚠️ Commercial deployment: Much of the published literature on airborne eDNA comes from academic proof-of-concept studies where deployment logistics are managed by PhD students or field researchers. Translating these methods to commercial monitoring programmes run by paid workers changes the cost-benefit picture substantially. Deploying, retrieving, cleaning, and processing large numbers of samplers carries real operational and financial implications that academic publications rarely quantify.
2.1 Active sampling
Active samplers use a powered mechanism — typically a fan, impeller, or pump — to draw a known volume of air through a filter or into a collection liquid at a controlled flow rate. The volume of air sampled is measurable, and sampling duration can be precisely controlled. This makes active sampling more amenable to quantitative comparisons between sites and time points.
Key characteristics of active samplers:
- High-volume stationary samplersTypically fixed units 500–1,500 m³ per hour; typically used for environmental monitoring; demonstrated to capture vertebrate eDNA (Littlefair et al. 2023) (e.g. Staplex, PM10 monitors)
- Medium-volume portable samplers Usually these include more professional setups with 50–300 L per minute; widely used in airborne microbiology and pollen monitoring (e.g. Coriolis Micro, Burkard Multi-Vial Cyclone):
- Low-volume DIY setupsMainly simple computer fans modified with filter holders; cost-effective for proof-of-concept work (Clare et al. 2022; Lynggaard et al. 2022). Some computer fans achieve surprisingly useful flow rates (Kroos et al. 2026).
- Liquid impingement samplersThese capture particles into liquid and may sustain higher volumetric flow rates and potentially preserve DNA better but have limiting deployment duration due to evaporation (e.g. BioSampler).
- Cyclone and Coriolis samplersThese use centrifugal force and air usually enters a cylindrical chamber tangentially, spiralling downward; particles above a threshold size are thrown to the outer wall and collected in a liquid reservoir at the base..
- Impactors and sticky-tape samplersCascade impactors direct airflow over a series of collection stages at progressively smaller nozzle sizes, depositing particles onto collection substrates by inertia in a size-resolved manner or on sticky tape or greased surfaces.
Comparison studies have found considerable variation among active sampler types even when sampling the same environment simultaneously. Polling et al. (2024) found that the Burkard spore trap — deployed continuously for 24 hours per day over one week in a Dutch field study — detected all vertebrate species recorded by co-located camera traps, plus additional taxa not captured by cameras. Berelson et al. (2025), reviewing sampler comparisons, note that such differences in detection performance between sampler types are consistent with differences in particle size capture efficiency. The ideal sampler would capture 100% of particles above a defined size threshold, but no such device currently exists.
Filter material and pore size are as consequential as sampler type. Kroos et al. (2026) found that coarse G4-rated fibreglass filters (capturing particles >10 µm) significantly outperformed finer F7-rated filters (1–10 µm) for targeted wallaby detection, recovering substantially more DNA — the coarser material appears to both capture a broader range of vertebrate-shed particle sizes and release DNA more readily during extraction. Bodawatta et al. (2025) independently reached the same conclusion in a multi-sampler comparison, finding that filters capable of retaining larger particles were more effective for characterising terrestrial vertebrate communities, and that larger filter surface area further improved vertebrate DNA detectability.
Active sampling is better suited to:
- Short-term, intensive surveys with precise temporal resolution
- Environments with low air movement (caves, dense forest understory)
- Studies requiring quantitative comparisons of DNA yield
- Biosecurity and pathogen surveillance applications
2.2 Passive sampling
Passive samplers do not use powered air movement. Instead, they rely on the natural deposition of airborne particles onto a collecting surface — through gravitational settling, electrostatic attraction, or impaction. Deployment is simple, requires no power, and samplers can remain in the field for days to weeks unattended. However, extended passive deployment introduces practical risks that are often underappreciated: DNA degrades continuously during deployment, particularly in warm, humid, or high-UV conditions, so longer deployment periods do not always yield proportionally more detectable signal.
Key passive sampling substrates demonstrated in the literature:
Passive sampling is better suited to:
- Long-term monitoring at sites where continuous power is unavailable.
- Citizen science applications and broad spatial coverage
- Landscape-scale biodiversity assessment (presence/absence focus).
2.3 Active vs passive: what the direct comparisons actually show
The intuitive expectation is that active samplers — which draw a controlled, known volume of air through the filter — should outperform passive ones simply by processing more air per unit time. The published comparison data are more complicated.
Jager et al. (2025) compared passive and active samplers simultaneously at Rotterdam Zoo, and the passive Nutshell DNAir sampler surpassed both active sampler types in species richness accumulation within 48 hours, continuing to accumulate new detections beyond 96 hours. It detected signal at the greatest distance (515 m from source animals) and showed a positive correlation with total species biomass. At short range and short duration, the active samplers yielded more DNA per unit time; over longer deployments, the passive sampler accumulated a more complete picture.
Lynggaard et al. (2024) deployed two active samplers of different flow rates side by side in a Danish mixed forest — a high-flow and a lower-flow unit — and found that neither consistently outperformed the other in terms of taxa detected. They concluded that successful detection appeared to depend more on sampler position and local air currents than on the volume of air processed or sampling duration.
Kroos et al. (2026), working in open farmland with targeted qPCR for invasive wallabies, found that at short distances (10–100 m), active samplers detected more reliably. At 1 km, detection rates for active (4.4%) and passive (6.7%) had converged — both methods detecting in only a small fraction of trials, largely dependent on whether a wind-carried particle happened to land on or pass through the sampler during the collection period.
Lin et al. (2025) confirmed that simple passive collectors were sufficient to detect considerable plant and vertebrate biodiversity in Chinese natural environments, finding results comparable to more expensive active systems.
What this body of evidence suggests is that the active vs passive question may be somewhat less important than it first appears — at least for the low- to medium-volume active systems typically used in published comparisons. The DNA flux from environmental targets is intermittent and spatially heterogeneous: airborne DNA arrives at a sampler in pulses shaped by wind, turbulence, and the stochastic movement of source animals. Whether a battery-powered pump processes 50 or 200 L/min may matter far less than whether the sampler happened to be in the right place during a pulse of signal.
As total sampled air volume increases substantially through longer high flow sampling or much higher flow rates the probability of encountering enough signal pulses to produce consistent detections improves considerably. Possibly the comparison studies published to date have mostly operated below this threshold.
2.4 Existing infrastructure: air quality monitors and radionuclide stations
A third category is often mentioned in the research sense — it is the repurposing of existing infrastructure. National air quality monitoring networks already deploy high-volume active samplers continuously, collecting particulate matter (PM10, PM2.5) onto filters for pollution analysis. These filters contain eDNA as a byproduct.
Littlefair et al. (2023) demonstrated this for UK air quality monitoring stations. Tournayre et al. (2025) scaled this to a national biodiversity survey. Sullivan et al. (2025) used 34 years of archived radionuclide monitoring filters — originally collected to detect radioactive fallout — to reconstruct ecosystem biodiversity back to 1974.
The strategic opportunity is real but should be qualified: existing air quality networks were designed for pollution monitoring at a regional scale, and their spatial density is modest relative to what fine-grained ecological surveillance would require. Nevertheless, these networks are already collecting material that contains recoverable biodiversity information, and retroactive analysis of archived filters requires no new field infrastructure.
2.5 Mobile Sampling
A fourth mode of collection — mobile sampling — mounts active or passive samplers on moving platforms such as vehicles, drones, or aircraft to survey large areas in a single deployment. Rather than waiting for DNA-laden particles to arrive at a fixed point, the sampler traverses the environment and accumulates signal from multiple source areas along the route. Métris & Métris (2023) described an aircraft-based approach capable of covering continental distances, demonstrating that airborne eDNA can be recovered from air collected at altitude and at speed. For ecological applications at landscape scale, vehicle-mounted samplers are more operationally tractable: Martino et al. (2025) compared stationary passive and vehicle-mounted filters across woodland and agricultural transects in Australia, detecting 49 vertebrate taxa overall, with five taxa recovered exclusively from the mobile method. They concluded the car mounted eDNA sampling would be very efficient for large scale rapid biomonitoring efforts.
The trade-offs are real: mobile sampling provides spatial breadth but sacrifices the detailed spatial accuracy of fixed stations, and protocols are not yet standardised. Possibly for monitoring programme design, the most practical approach may be to use mobile surveys as a large scale discovery tool to identify areas of interest, then deploy stationary samplers at key locations.
3. DNA extraction from air filters
Once a filter has been collected, extracting sufficient DNA for downstream analysis is a critical step. Air filters present specific challenges not encountered in aquatic eDNA work: the total amount of biological material may be small, it is distributed across a large filter area, and the physical matrix of the filter itself can inhibit extraction.
Clare et al. (2022) developed a protocol adapted from tissue extraction kits (Qiagen DNeasy Blood & Tissue), increasing buffer volumes to ensure filter submersion, using QIA Shredder columns to process remaining filter material, and extending lysis times overnight at 56°C with shaking. This protocol has been widely adopted and modified by subsequent groups.
Key considerations in extraction:
4. Sequencing strategies: PCR, metabarcoding vs shotgun
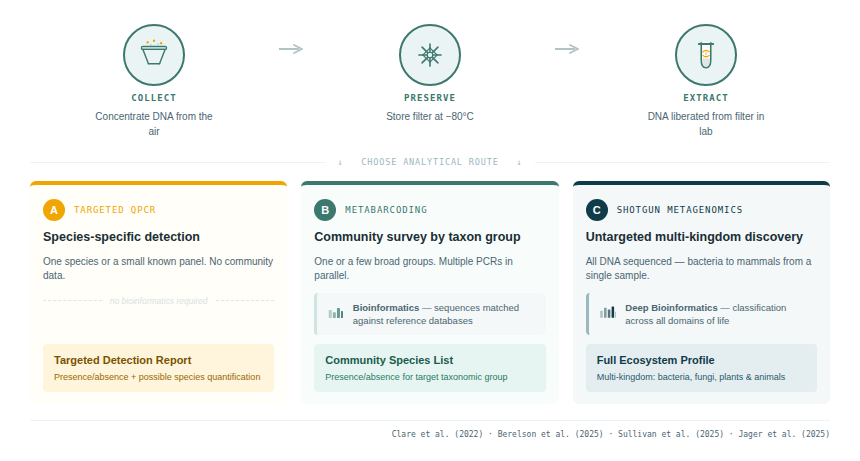
Once DNA has been extracted, three broad analytical strategies are available, each with distinct advantages and limitations. The choice between them depends on whether the objective is targeted detection of a known species, community-wide surveillance of a taxonomic group, or untargeted discovery across the tree of life.
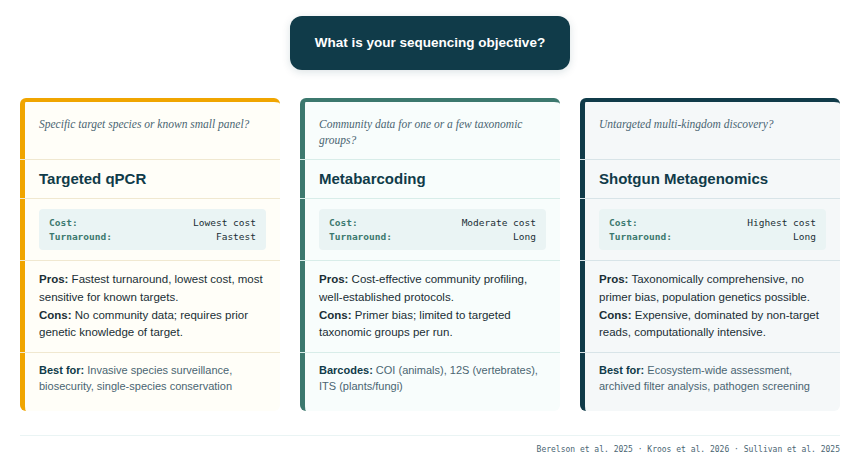

4.1 Targeted PCR and qPCR
The simplest and most sensitive approach is targeted PCR or quantitative PCR (qPCR), using primers and probes designed specifically for a single species or a small number of known targets. This approach is not a sequencing strategy per se — it typically produces a presence/absence or abundance signal rather than sequence data — but it is the most operationally tractable option for biosecurity surveillance, invasive species monitoring, or single-species conservation applications.
Targeted PCR is better suited to:
- Single-species or small-panel surveillance (invasive species, pathogens, endangered species)
- Regulatory or biosecurity applications requiring legally defensible, species-specific data
- Remote or low-resource settings where sequencing infrastructure is unavailable
4.2 DNA metabarcoding (amplicon sequencing)
Metabarcoding uses PCR to amplify a specific short DNA region — a "barcode" — using primers that target a broad taxonomic group. Many different target regions exist, but the most commonly used barcodes include:
This approach is highly sensitive, cost-effective, and well-suited to detecting rare taxa. The primary limitation is taxonomic scope: a single PCR reaction targets one broad group (e.g. vertebrates), and multiple reactions must be run in parallel to capture multi-taxa diversity. Additionally, primer bias can cause some taxa to be amplified more efficiently than others, distorting the apparent community composition (Alberdi et al. 2018).
Total air volume processed is a key sensitivity driver across all three analytical approaches — targeted qPCR, metabarcoding, and shotgun sequencing alike. More air means more DNA molecules on the filter, directly lowering the detection threshold for rare or distant taxa. Academic portable active samplers typically run at 100–300 L/min; over a 4–8 hour field deployment that yields roughly 25–150 m³ per sample. High-volume station-based systems (500–1,500 L/min, equivalent to PM10 air quality monitors) running for 24 hours collect on the order of 700–2,200 m³. Generally, higher processed volumes seem to improve sensitivity but sometime require correspondingly more complex extraction to recover the DNA from larger filter areas.
4.3 Shotgun metagenomics
Shotgun sequencing does not use targeted amplification. Instead, all DNA in a sample — from all organisms simultaneously — is fragmented, sequenced, and then computationally sorted into taxonomic bins. This approach is taxonomically comprehensive and not subject to primer bias, but it generates enormous datasets dominated by non-target DNA (environmental background, bacteria, non-biological contaminants).
Sullivan et al. (2025) used deep shotgun sequencing to reconstruct 34 years of ecosystem biodiversity from archived filters, detecting over 2,700 genera across all domains of life in a single analytical framework. Nousias et al. (2025) demonstrated that shotgun sequencing of air samples can characterise whole biomes, detect population genetic structure, and screen for pathogens simultaneously. The cost-effectiveness of shotgun sequencing has improved dramatically, but it remains more expensive per sample than targeted metabarcoding, particularly for routine monitoring applications.
4.4 Nanopore sequencing
Oxford Nanopore's MinION device produces long reads (thousands to tens of thousands of base pairs) and can operate portably in the field. Berelson et al. (2025) describe this as an emerging option for airborne eDNA, particularly for pathogen surveillance where real-time results are needed. Long reads improve taxonomic resolution for difficult groups and can capture structural variants. Current limitations include a higher error rate than short-read platforms and the need for specialist computational skills for assembly.
4.5 Enhancing signal quality
Because airborne eDNA samples often contain low template concentrations, several steps can improve the reliability of results beyond simply choosing the right sequencing strategy.
PCR replication is standard good practice: running multiple independent PCR reactions per sample (typically two to three) and pooling them before sequencing increases detection probability for rare taxa and helps identify artefacts — a species appearing consistently across replicates is more credible than one appearing in just one. Most published studies pool replicates before library preparation for cost efficiency; keeping replicates as separate sequencing libraries provides stronger consistency filtering but is more expensive.
Inhibition management and extraction chemistry matter considerably in airborne eDNA. Filters can carry co-extracted compounds (humic material, pigments, particulate matter) that inhibit PCR. Diluting extracts, using inhibitor-removal columns, or adjusting extraction chemistry can recover signal from filters that would otherwise fail amplification. Laboratories have developed different preferred approaches to this, and there is not yet a single standard method.
Blocking oligonucleotides are used in some protocols to suppress dominant non-target signals — most commonly human DNA in 12S vertebrate metabarcoding. Where such blocking steps are applied they can substantially improve the wildlife signal-to-noise ratio, but their use and optimisation varies between laboratories.
5. Bioinformatic analysis and reference databases
Sequencing produces raw data — millions of short DNA sequences. Converting these into a species list requires bioinformatic processing, which includes quality filtering, chimera removal, clustering into amplicon sequence variants (ASVs) or operational taxonomic units (OTUs), and taxonomic assignment against a reference database.
The quality of taxonomic assignment is directly limited by the completeness of available reference databases. For vertebrates, databases such as NCBI GenBank and the Barcode of Life Data System (BOLD) provide reasonable coverage of described species in developed countries. For plants, insects, and microorganisms in tropical or poorly surveyed regions, coverage is sparse. Sullivan et al. (2025) found that 76% of shotgun sequencing reads from a Swedish boreal forest archive could not be classified to a known organism. This figure requires careful interpretation: it reflects a combination of genuinely uncharacterised taxa, short or degraded fragments that resist classification regardless of database completeness, and the inherent noise in deep shotgun sequencing of environmental material. It does not straightforwardly mean that 76% of additional species would be identifiable if databases were better.
Reference database gaps are a real constraint, but their policy significance is often overstated. For most taxa that matter most to government users — endangered species, invasive species, quarantine organisms, and commercially important vertebrates — reference sequences are well represented in databases, particularly in the Western world. The species least likely to be in reference databases tend to be those in remote tropical regions where airborne eDNA would in any case face the greatest deployment challenges. As global reference databases expand through initiatives such as the Earth BioGenome Project and national DNA barcoding programmes, the same archived air filters yield additional value through two distinct routes: re-running the bioinformatic matching of existing sequence data against an expanded reference database (no new laboratory work required), or physically re-extracting and re-sequencing the stored filter using improved extraction or sequencing methods (warranted when library preparation or sequencing chemistry has advanced substantially). The former is computationally inexpensive and immediately applicable to any study that has retained its raw sequence files; the latter recovers signal lost to earlier technical limitations but requires the filter material to have been adequately preserved.
6. Caveats and open questions
No universal standard exists. Different sampler types yield systematically different results. Without standardised protocols, data from different studies cannot be reliably compared. The UK NPL/LGC collaboration (NPL/LGC 2025) and the Southern eDNA Society workshop (Tulloch et al. 2025) both identified standardisation as the most urgent priority for the field.
Temporal dynamics are poorly understood. How rapidly does the eDNA signal change? Lynggaard et al. (2024) showed weather-dependent variation in vertebrate detections over three days in a Danish forest. Sullivan et al. (2025) showed that weekly filters capture seasonal patterns. But the minimum sampling duration needed to reliably characterise a community is unknown and likely context-dependent.
Power and logistics constrain active sampler deployment in some settings. Active samplers require electricity, which limits remote deployment to sites with battery packs, vehicle power, or solar charging. Passive samplers avoid this constraint but trade off temporal precision and sensitivity for rare taxa. Field-deployed samplers of any type are also vulnerable to tampering, theft, and vandalism in accessible locations — a practical factor that should be factored into monitoring programme design.
References
- Berelson MFG et al. (2025). From air to insight: the evolution of airborne DNA sequencing technologies. Microbiology 171:001564. https://doi.org/10.1099/mic.0.001564
- Clare EL et al. (2022). Measuring biodiversity from DNA in the air. Current Biology 32(3):693–700. https://doi.org/10.1016/j.cub.2021.11.064
- Jager H et al. (2025). A breath of fresh air: comparative evaluation of passive versus active airborne eDNA sampling strategies. bioRxiv. https://doi.org/10.1101/2025.03.26.645491
- Johnson M & Barnes MA (2024). Macrobial airborne environmental DNA analysis: A review. Molecular Ecology Resources 24:e13998. https://doi.org/10.1111/1755-0998.13998
- Lin R et al. (2025). Robust passive sampling of airborne environmental DNA to monitor plants and animals. Methods in Ecology and Evolution. https://doi.org/10.1111/2041-210X.70102
- Littlefair JE et al. (2023). Air-quality networks collect environmental DNA. Current Biology 33:R426–R428. https://doi.org/10.1016/j.cub.2023.04.036
- Lynggaard C et al. (2024). Airborne environmental DNA captures terrestrial vertebrate diversity in nature. Molecular Ecology Resources 24:e13840. https://doi.org/10.1111/1755-0998.13840
- Newton JP et al. (2024). Spider webs capture environmental DNA from terrestrial vertebrates. iScience 27:108904. https://doi.org/10.1016/j.isci.2024.108904
- Nousias O et al. (2025). Shotgun sequencing of airborne eDNA achieves rapid assessment of whole biomes. Nature Ecology & Evolution. https://doi.org/10.1038/s41559-025-02711-w
- Sanders NJ et al. (2023). Exploring the use of dust traps as passive samplers for airborne eDNA. Environmental DNA 5:e392. https://doi.org/10.1002/edn3.392
- Sullivan AR et al. (2025). Airborne eDNA captures three decades of ecosystem biodiversity. Nature Communications 16:11281. https://doi.org/10.1038/s41467-025-67676-7
- Tournayre O et al. (2025). First national survey of terrestrial biodiversity using airborne eDNA. Scientific Reports 15:19247. https://doi.org/10.1038/s41598-025-03650-z
- Tulloch RL et al. (2025). Winds of Change: Charting a Pathway to Ecosystem Monitoring Using Airborne Environmental DNA. Environmental DNA. https://doi.org/10.1002/edn3.70134
- Alberdi A et al. (2018). Scrutinizing key steps for reliable metabarcoding of environmental samples. Methods in Ecology and Evolution 9:134–147. https://doi.org/10.1111/2041-210x.12849
- Gygax D et al. (2026). Monitoring terrestrial vertebrates with airborne DNA in the Luangwa Valley, Zambia. bioRxiv 2026.03.11.711018. https://doi.org/10.64898/2026.03.11.711018
- Bodawatta KN et al. (2025). Advanced airborne eDNA sampling allows robust spatiotemporal characterisation of vertebrate communities. Research Square preprint. https://doi.org/10.21203/rs.3.rs-5693469/v1
- Kroos GC et al. (2026). Targeted airborne eDNA detection of pest wallabies: effects of sampler type and distance. bioRxiv 2025.10.26.684691. https://doi.org/10.1101/2025.10.26.684691
- NPL/LGC (2025). Standardising airborne eDNA for biodiversity monitoring: a UK collaboration to advance measurement science. National Physical Laboratory / National Measurement Laboratory at LGC. https://www.npl.co.uk/news/uk-launches-initiative-to-advance-measurement-science-for-airborne-edna-for-biodiversity-monitoring
- Martino F et al. (2025). eDNA on the go: a direct comparison of fixed and vehicle mounted airborne eDNA sampling methods for terrestrial vertebrate species detection at large spatial scales. bioRxiv 2025.09.02.673667. https://doi.org/10.1101/2025.09.02.673667
- Métris KL, Métris J (2023). Aircraft surveys for air eDNA: probing biodiversity in the sky. PeerJ 11:e15171. https://doi.org/10.7717/peerj.15171
- Elbrecht V & Leese F (2017). Validation and development of COI metabarcoding primers for freshwater macroinvertebrate bioassessment. Frontiers in Environmental Science 5:11. https://doi.org/10.3389/fenvs.2017.00011
- Klepke MJ et al. (2022). Accumulation and diversity of airborne, eukaryotic environmental DNA. Environmental DNA 4:1323–1339. https://doi.org/10.1002/edn3.340
- Polling M et al. (2024). Continuous daily sampling of airborne eDNA detects all vertebrate species identified by camera traps. Environmental DNA 6(4):e591. https://doi.org/10.1002/edn3.591