Short Answer: The range of organisms detectable from airborne eDNA has grown dramatically since the first zoo studies in 2022. Microbes, fungi, plants, insects and vertebrates have all been successfully detected. The most comprehensive study to date — Sullivan et al. (2025) — identified over 2,700 genera spanning every domain of life from a single archive of archived aerosol filters. In a national UK survey, Tournayre et al. (2025) recovered over 1,100 taxa including confirmed invasive species not previously recorded at those sites. The honest answer is: probably almost everything, subject to the completeness of reference databases.
1. The breadth of detection: an expanding catalogue
The short answer is almost everything — subject to reference database completeness and the evidence maturity for each group.
The history of airborne eDNA detection is, in large part, the history of progressively expanding the catalogue of what can be found. Each major study has pushed the boundaries of what is considered detectable.
The first vertebrate zoo studies (Clare et al. 2022; Lynggaard et al. 2022) detected mammals and birds. Roger et al. (2022) extended this to arthropods. Johnson et al. (2021) showed plants could be detected beyond their pollen season. Abrego et al. (2024) demonstrated global spatial and seasonal patterns of fungal diversity from airborne samples. And Sullivan et al. (2025) collapsed all these categories simultaneously, detecting organisms from bacteria to mammals in a single non-targeted framework.
It is now accurate to say that airborne eDNA is, in principle, a whole-ecosystem surveillance method — not a single-taxon detection tool.
2. Vertebrates
2.1 Mammals and birds
Mammals and birds are the best-studied taxon group in airborne eDNA. Unlike plants (which release pollen) or fungi (which actively discharge spores), vertebrates have no dedicated airborne dispersal stage — they shed detectable quantities of DNA simply by existing in an environment: breathing, moving, excreting, and through grooming, scratching, and predator–prey interactions that generate fine-particle biological material. This means detection does not depend on a particular season or behaviour — the signal is continuous wherever animals are present.
Mammals and birds are the best-studied taxon group in airborne eDNA, partly because their detection was the most surprising — and partly because the two simultaneous 2022 zoo studies provided immediate, highly visible evidence that the method worked.
Clare et al. (2022) detected 25 species at Hamerton Zoo UK, including tigers, lemurs, dingoes, and — critically — wild species outside the zoo perimeter: Eurasian hedgehog (endangered in the UK) and muntjac deer. Lynggaard et al. (2022) detected 49 species at Copenhagen Zoo from just 40 samples, including mammals, birds, amphibians, reptiles and fish. Both teams found that detection probability increased with proximity to source animals and with biomass, but detection was possible at hundreds of metres from source.
Subsequent studies moved from zoo settings to wild landscapes. Lynggaard et al. (2024) collected 143 air samples over three days in a mixed Danish forest and detected 64 bird, mammal, fish and amphibian taxa, representing over a quarter of all terrestrial vertebrates known to occur in the region. Polling et al. (2024) compared airborne eDNA against camera trap records across three Netherlands habitats over three weeks, detecting all vertebrate species recorded by camera traps plus additional taxa the cameras missed entirely.
Warmer et al. (2025) conducted the first direct three-way comparison of airborne eDNA, manual surveys, and acoustic monitoring in a Dutch agroforest. Airborne eDNA and acoustic monitoring both detected a large proportion of species found by traditional methods, plus additional species undetected by the other approaches. Airborne eDNA performed particularly well for nocturnal and cryptic species.
2.2 Amphibians, reptiles and fish
Among the most striking findings of the early zoo studies was the detection of fish, amphibians and reptiles from air samples. These are not typically thought of as organisms that shed airborne DNA. The most likely explanation is that their DNA is present in the air via indirect routes: insect prey carrying DNA from their gut contents, aerosols from aquatic habitats, or particulate matter transported by wind from nearby water bodies.
O'Donnell et al. (2025) systematically investigated water-to-air eDNA transfer, demonstrating for the first time under field conditions that aquatic organisms (in their case, spawning Coho salmon) can be consistently detected from passive air samples collected above a stream. This cross-medium transfer has significant implications for the interpretation of airborne eDNA signals and potentially for freshwater monitoring.
2.3 Avian diversity at continental scale
Birds deserve special mention because their migratory nature makes continental-scale monitoring both particularly challenging and particularly valuable. Sullivan et al. (2025) found that bird eDNA trends reconstructed from their Swedish archive correlated strongly with independent point-transect survey data (adjusted R² = 0.60), providing the first multi-decade validation of airborne eDNA as a reliable indicator of vertebrate abundance trends.
3. Insects and invertebrates
Insects represent perhaps the most strategically important taxon group for airborne eDNA, given the scale and speed of the global insect decline crisis — and the critical role insects play in ecosystem functioning and agricultural productivity, from pollination and nutrient cycling to pest regulation and food web support. Hallmann et al. (2017) documented more than 75% decline in flying insect biomass over 27 years in protected German areas — and this is one of the better-monitored regions globally. For most of the world, insect population trends are essentially unknown.
Pumkaeo et al. (2021) were among the first to demonstrate detection of insect bioaerosols from air, establishing that insect-derived DNA can be recovered from atmospheric samples. Roger et al. (2022) then provided a rigorous proof of concept for arthropod detection via airborne eDNA metabarcoding in natural open-field conditions, recovering a diverse insect community including species not observed by concurrent visual surveys. Both studies contributed to establishing insects as detectable via airborne eDNA, and multiple subsequent field studies have confirmed and expanded on these findings.
The particular value of airborne eDNA for insects is that it detects nocturnal, cryptic and small-bodied species that conventional monitoring systematically underrepresents. As Warmer et al. (2025) noted, airborne eDNA detected a large proportion of birds and mammals found by traditional methods, plus a range of species undetected by the latter — and this complementarity was especially strong for species with small bodies or nocturnal activity patterns.
4. Fungi
Fungi are among the most abundant biological particles in the near-surface atmosphere, and their detection via airborne DNA methods has a longer history than macroorganism eDNA. The Global Spore Sampling Project (Ovaskainen et al. 2024) has now produced standardised fungal eDNA data from sites on every inhabited continent, making it the most geographically comprehensive airborne eDNA dataset in existence.
Abrego et al. (2024), in a landmark Nature paper, used global airborne spore sampling to demonstrate that fungal diversity follows predictable spatial and seasonal patterns linked to vegetation type, climate zone and land use. This is the first demonstration that airborne eDNA can be used to track ecosystem-level changes in fungal communities at global scale.
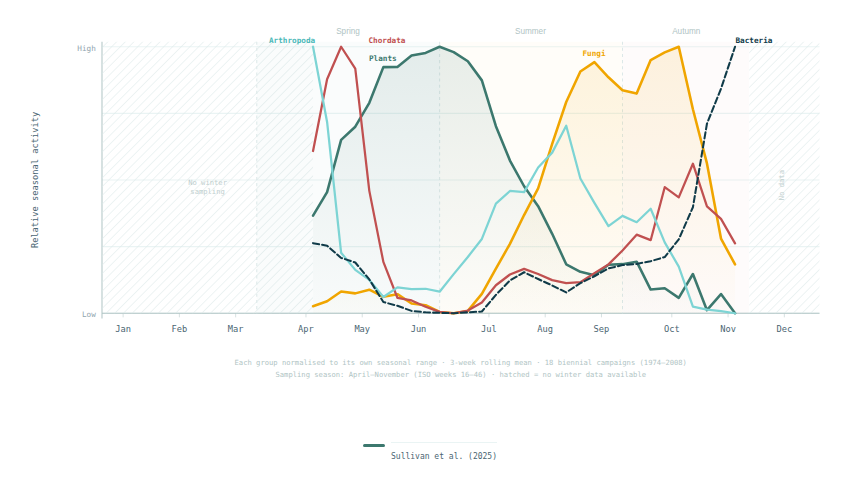
Sullivan et al. (2025) found that fungal eDNA showed a characteristic bimodal seasonality in their Swedish boreal archive, with peaks slightly in spring and massively in autumn driven by fruiting body sporulation cycles — distinct from plant patterns and consistent with decades of aerobiology literature on fungal spore calendars.
5. Plants
5.1 Beyond pollen: whole-plant community detection
Plants have the longest history of airborne DNA detection — the monitoring of pollen via volumetric spore traps has been standard practice in aerobiology since Hirst (1952). What is new is the detection of non-pollen plant DNA and the extension to whole-community metabarcoding.
Johnson et al. (2021) compared airborne eDNA metabarcoding against a traditional plant community survey and found that the eDNA method detected more diversity with less sampling effort. Johnson et al. (2019) demonstrated that even entomophilous species — plants that rely on insects rather than wind for pollination, and therefore produce minimal airborne pollen — can be detected via airborne eDNA. This suggests that plant DNA enters the air through leaf fragment release, cuticle particles and other non-pollen pathways.
Lin et al. (2025) confirmed that passive eDNA sampling effectively captures plant communities in natural environments, finding that prolonged sampling integrates community composition over time rather than instantaneous snapshots.
Tournayre et al. (2025), in their UK national survey, identified plants across a wide range of life history traits — from common agricultural weeds to native woodland species — suggesting that airborne eDNA captures a broadly representative cross-section of local plant communities.
5.2 Seasonal dynamics
Plant eDNA signals in air are not constant. They peak during reproductive periods (particularly for wind-pollinated species) and vary with weather conditions. Sullivan et al. (2025) found that most plant eukaryotes showed a single seasonal peak, with grasses peaking in summer and trees showing an early spring peak driven by flowering. This means that study design must account for seasonality: a single air sample taken in one season may not represent annual plant diversity. This parallels well-established aerobiology findings for pollen (Sofiev et al. 2013; Mandrioli et al. 2001).
6. Microorganisms: bacteria, archaea and viruses
Microbial aerobiology predates macroorganism airborne eDNA by decades. Atmospheric microbiome research has established that bacterial diversity in outdoor air is high, seasonally variable, and connected to terrestrial sources including soil, vegetation and human activity (Bowers et al. 2011; Caliz et al. 2018; de Groot et al. 2021).
In the context of macroorganism airborne eDNA studies, microorganisms often dominate the sequencing data. Sullivan et al. (2025) found that prokaryotic genera showed a bimodal seasonal pattern with peaks in spring and autumn — contrasting with the single-season peak of most eukaryotes. This bimodal pattern was unexpected and differed from some earlier aerobiology literature, suggesting it may be a feature specific to boreal ecosystems or to the high-latitude site in northern Sweden. The potential applications of microbial airborne eDNA for public health — including pathogen surveillance, antimicrobial resistance monitoring, and early warning of disease outbreaks — are significant and the subject of active research.
7. The reference database ceiling
The best understood constraint on what can be detected is not the sensitivity of the method — it is the completeness of taxonomic reference databases. Sullivan et al. (2025) found that 76% of sequencing reads from a biologically rich boreal ecosystem could not be assigned to any known taxon. This figure requires careful interpretation: it reflects a combination of genuinely uncharacterised taxa, short or degraded fragments that resist classification regardless of database completeness, and the inherent noise of deep shotgun sequencing of environmental material. It does not straightforwardly mean that those organisms are absent, nor that they would all be identifiable if databases were better.
This has a positive implication that is often overlooked: the same archived air filters can be re-analysed as databases improve. The biological samples do not need to be recollected. As national barcoding initiatives, the Earth BioGenome Project, and taxonomic digitisation efforts expand global reference coverage over the coming decade, retrospective re-analysis of archived filters will progressively reveal the biodiversity they already contain.
8. What airborne eDNA detects that other methods miss
A consistent finding across comparative studies is that airborne eDNA is complementary to, not a replacement for, other monitoring methods. Each approach has blind spots that others fill.
| Method | Strengths | Misses |
|---|---|---|
| Airborne eDNA | Cryptic, nocturnal, small-bodied taxa; multi-taxon simultaneously | Quantification; fine spatial resolution |
| Camera traps | Behavioural context; images; species confirmation | Small-bodied; nocturnal (partial); invertebrates |
| Acoustic monitoring | Real-time; directional; temporal resolution | Non-vocalising taxa |
| Visual transects | Species confirmation; abundance | Cryptic; nocturnal; invertebrates |
| Citizen science | Scale; phenology | Charismatic species bias; inconsistent effort |
Tournayre et al. (2025) found that citizen science (iNaturalist) detected more taxa overall than airborne eDNA, but eDNA was substantially better at detecting taxa that are "harder to spot or identify visually" — precisely the species most at risk of being undercounted in biodiversity assessments. It is worth noting that citizen science coverage varies considerably by country: the UK has an exceptionally rich and long-established network of volunteer naturalists, making iNaturalist comparisons there unusually favourable. In countries with less developed citizen science infrastructure, the comparative advantage of airborne eDNA over volunteer recording is likely to be considerably greater.
References
- Abrego N et al. (2024). Airborne DNA reveals predictable spatial and seasonal dynamics of fungi. Nature 631(8022):835–842. https://doi.org/10.1038/s41586-024-07658-9
- Bowers RM et al. (2011). Sources of bacteria in outdoor air across cities in the midwestern United States. Applied and Environmental Microbiology 77:6350–6356. https://doi.org/10.1128/AEM.05498-11 [UNVERIFIED — confirm DOI]
- Clare EL et al. (2022). Measuring biodiversity from DNA in the air. Current Biology 32(3):693–700. https://doi.org/10.1016/j.cub.2021.11.064
- de Groot GA et al. (2021). Detection of plant and animal biodiversity using airborne environmental DNA. Environment International 158:106551. https://doi.org/10.1016/j.envint.2021.106551
- Hallmann CA et al. (2017). More than 75 percent decline over 27 years in total flying insect biomass. PLOS ONE 12:e0185809. https://doi.org/10.1371/journal.pone.0185809
- Johnson MD & Barnes MA (2019). The detection of a non-anemophilous plant species using airborne eDNA. PLoS ONE 14:e0225262. https://doi.org/10.1371/journal.pone.0225262
- Johnson MD et al. (2021). Airborne environmental DNA metabarcoding detects more diversity, with less sampling effort, than a traditional plant community survey. BMC Ecology and Evolution 21:218. https://doi.org/10.1186/s12862-021-01947-x
- Lin R et al. (2025). Robust passive sampling of airborne environmental DNA to monitor plants and animals. Methods in Ecology and Evolution. https://doi.org/10.1111/2041-210X.70102
- Lynggaard C et al. (2022). Airborne environmental DNA for terrestrial vertebrate community monitoring. Current Biology 32(3):701–707. https://doi.org/10.1016/j.cub.2021.12.014
- Lynggaard C et al. (2024). Airborne environmental DNA captures terrestrial vertebrate diversity in nature. Molecular Ecology Resources 24:e13840. https://doi.org/10.1111/1755-0998.13840
- Mandrioli P et al. (2001). Aerobiology: past, present and future. Aerobiologia 17:1–3. https://doi.org/10.1023/A:1007602928928
- Nousias O et al. (2025). Shotgun sequencing of airborne eDNA achieves rapid assessment of whole biomes. Nature Ecology & Evolution. https://doi.org/10.1038/s41559-025-02711-w
- O'Donnell S et al. (2025). Passive air sampling detects environmental DNA transfer from water into air. Scientific Reports. https://doi.org/10.1038/s41598-025-26293-6
- Ovaskainen O et al. (2024). Global Spore Sampling Project: A global, standardized dataset of airborne fungal DNA. Scientific Data 11:583. https://doi.org/10.1038/s41597-024-03410-0
- Polling M et al. (2024). Continuous daily sampling of airborne eDNA detects all vertebrate species identified by camera traps. Environmental DNA 6:e591. https://doi.org/10.1002/edn3.591
- Pumkaeo P et al. (2021). Detection and monitoring of insect traces in bioaerosols. PeerJ 9:e10862. https://doi.org/10.7717/peerj.10862
- Roger F et al. (2022). Airborne environmental DNA metabarcoding for the monitoring of terrestrial insects. Environmental DNA 4:790–807. https://doi.org/10.1002/edn3.290
- Sullivan AR et al. (2025). Airborne eDNA captures three decades of ecosystem biodiversity. Nature Communications 16:11281. https://doi.org/10.1038/s41467-025-67676-7
- Tournayre O et al. (2025). First national survey of terrestrial biodiversity using airborne eDNA. Scientific Reports 15:19247. https://doi.org/10.1038/s41598-025-03650-z
- Warmer J et al. (2025). Validating airborne eDNA using manual surveys, acoustic monitoring and camera traps. Environmental DNA. https://doi.org/10.1002/edn3.70222