Short Answer: Water eDNA revolutionised aquatic biodiversity monitoring. Soil eDNA is invaluable for below-ground communities. Airborne eDNA is the newest substrate, but not simply a terrestrial equivalent of water eDNA — it behaves differently in space and time, detects different taxa with different efficiencies, and captures landscape-scale information rather than point-source samples. The three substrates are most powerful when used in combination: each has blind spots the others fill.
1. The eDNA substrate landscape
Environmental DNA is not a single method — it is a family of approaches that share a common analytical core (DNA extraction and sequencing) but differ fundamentally in how genetic material accumulates, persists, and is sampled in different environmental matrices.
The three primary substrates are:
A fourth approach — sometimes termed forensic environmental sampling — collects deposited biological material from surfaces rather than from a fluid medium. This includes surface swabbing (e.g. vegetation surfaces, rocks, fences), faecal DNA collection, and targeted substrate sampling at points of known animal contact (urine marks, resting sites, den entrances). Forensic sampling is not a substrate in the same sense as air, water, or soil, but it is increasingly used alongside them for targeted single-species detection and attribution. It shares the analytical workflow of eDNA but differs in that the sampling is targeted rather than general.
Each substrate reflects a different ecological signal, operates at a different spatial and temporal scale, and is subject to different physical and chemical controls on DNA persistence and transport.
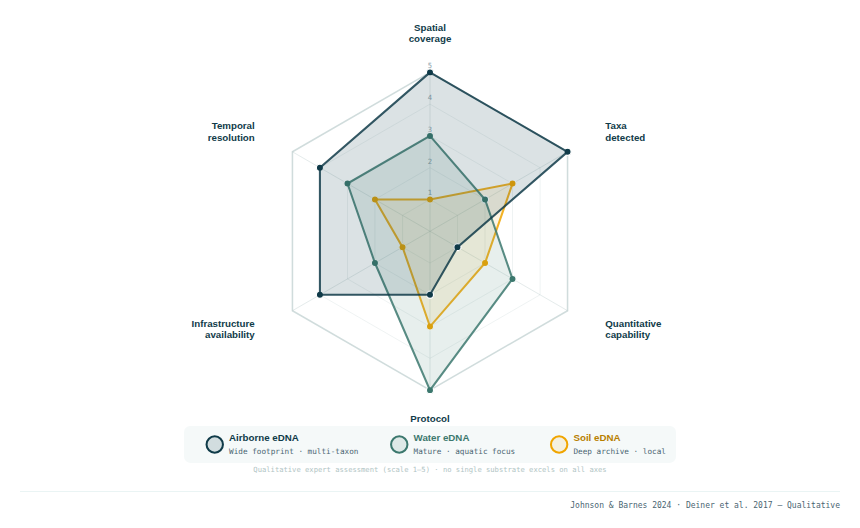
2. What each substrate captures
2.1 Water eDNA
Aquatic eDNA has been most extensively developed for fish, amphibians, and aquatic macroinvertebrates. The fundamental principle is that aquatic organisms shed DNA into the water column through mucus, scales, faeces, and cellular secretions. This DNA is detectable downstream of the source organism, with transport distances in flowing water typically ranging from hundreds of metres to a few kilometres before degradation or dilution makes detection unreliable (Jane et al. 2015; Shogren et al. 2017).
Strengths of water eDNA:
- Highly sensitive for aquatic vertebrates, particularly fish
- Excellent for detecting rare or invasive aquatic species (Ficetola et al. 2008; Rees et al. 2014)
- Established regulatory protocols exist (great crested newt in UK; Asian carp monitoring in USA)
- Quantitative frameworks are more developed than for airborne eDNA
Limitations:
- Primarily captures aquatic organisms; terrestrial organisms are detectable at water surfaces but less reliably than from air
- DNA degrades rapidly in some conditions (UV exposure, high temperature, microbial activity)
- Spatial resolution is limited by water movement — sample represents an upstream catchment of uncertain extent
- Logistically demanding in remote or difficult-to-access water bodies
2.2 Soil eDNA
Soil and sediment eDNA is dominated by plant and microbial communities or marine organism in marine settings. DNA persists in soil for much longer than in water or air — potentially thousands of years in frozen or anaerobic conditions (Willerslev et al. 2003). This makes sedimentary ancient DNA (sedaDNA) a powerful tool for reconstructing past biodiversity and tracking ecosystem changes over centuries (Parducci et al. 2017; Alsos et al. 2022).
Strengths of soil eDNA:
- Exceptional for plant community characterisation (both present and historical)
- Excellent temporal depth — sediment cores provide decade-to-millennium biodiversity archives
- Captures below-ground biodiversity (soil invertebrates, fungi, bacteria) that air and water miss
- Lower temporal variability than air or water — integrates over longer periods
Limitations:
- Limited spatial coverage per sample — a soil core represents centimetres to metres, not kilometres
- DNA from different time periods is mixed; temporal resolution requires careful stratigraphy
- Below-ground organisms dominate; flying or highly mobile species may be underrepresented
- Collection is labour-intensive at scale
2.3 Airborne eDNA
Airborne eDNA captures a fundamentally different cross-section of biodiversity. The key distinguishing features are its spatial coverage (a single sampler captures signal from a radius of kilometres), its multi-taxon breadth (bacteria to mammals in a single sample), and its direct relevance to the terrestrial world where human land use is most intense and where biodiversity monitoring decisions most frequently need to be made. Unlike aquatic eDNA, which is limited to organisms in or near water, airborne eDNA provides access to the full terrestrial biota — the landscapes of agriculture, urbanisation, and conservation management. How this is achieved in practice will involve a combination of dedicated samplers, opportunistic use of existing air quality infrastructure, and integration with other monitoring approaches.
Strengths of airborne eDNA:
- Taxonomically non-selective: detects microbes, plants, fungi, insects and vertebrates simultaneously
- Large spatial footprint: a single sampler represents a landscape-scale catchment
- Compatible with existing infrastructure: air quality filter networks already in operation globally
- Particularly effective for difficult-to-detect taxa: cryptic, nocturnal, small-bodied species
- Temporal continuity: continuous monitoring possible via networked stations or frequent sample collection and turnover
Limitations:
- DNA cannot be easily attributed to a precise source location
- Transport distance uncertainty complicates presence/absence interpretation
- Less developed protocols and regulatory frameworks than water eDNA
- Quantification is not yet reliable for most taxa
3. Direct comparisons: what the studies show
3.1 Air vs. water for aquatic organisms
Ip et al. (2025a) made the first systematic comparison of air and water sampling for the same aquatic target species (spawning Coho salmon in a Washington State stream). They found that DNA from the salmon was consistently detectable from passive air samples collected above the water surface, demonstrating that aquatic organisms can be detected from air through water-to-air transfer of eDNA via surface aerosols. Despite an approximately 25,000-fold dilution relative to the water signal, passive air collectors captured quantitative airborne eDNA signals that closely paralleled salmon counts over a six-week spawning period.
A follow-up study by the same group (Ip et al. 2025b; preprint) extended this approach to community-level vertebrate metabarcoding at two urban–wildland interface sites, collecting 27 paired air-and-water samples. They found substantial overlap in vertebrate species detected across both media: abundant aquatic species tracked temporally across air and water within 24 hours, while rare or low-abundance taxa were the first to drop out of cross-medium transfer. This finding has practical implications: it suggests that community-level signals do cross the air-water boundary, but detection sensitivity is strongly modulated by species abundance, meaning that airborne detection of aquatic organisms cannot replace in-water sampling for rare species.
Taken together, these results suggest that air monitoring near rivers and lakes may provide information about aquatic communities as a byproduct, and that the strict separation between "aquatic" and "terrestrial" eDNA sampling may be less absolute than previously assumed. They also raise a less discussed possibility — that airborne eDNA originating from one water body could be deposited in an adjacent one via atmospheric transport and wet or dry deposition. Until recently this was not recognised as a meaningful route of eDNA dispersal between water bodies (transfer via birds carrying eDNA on their bodies was acknowledged, but airborne particle transport was not). This has potential implications for the interpretation of aquatic eDNA signals in water bodies close to each other and for biosecurity protocols around invasive aquatic species.
3.2 Air vs. soil for microbial and fungal communities
The comparison between airborne and soil eDNA is most developed for microorganisms and fungi — the dominant signals in both matrices. Fungal communities are exceptionally well-represented in air via spore release and in soil via mycelial networks and spore banks; the two substrates capture overlapping but distinct fungal communities at different life-cycle stages (Abrego et al. 2024; Ovaskainen et al. 2024). Bacterial communities in air are strongly shaped by nearby soil surfaces, with terrestrial soil the dominant source of outdoor airborne bacteria (Bowers et al. 2011).
For plant communities specifically, the picture is more nuanced. Airborne eDNA integrates over a landscape-scale catchment and is strongest during reproductive periods; soil eDNA captures local, point-specific plant communities with high fidelity and longer temporal depth. Johnson et al. (2021) found that airborne eDNA metabarcoding detected more plant diversity than traditional field surveys, though direct systematic comparisons with soil eDNA for the same sites are limited. Detection of plant DNA from soil eDNA approaches is rarely the primary method of choice — it is incidental to microorganism-focused studies — and caution is warranted before treating airborne vs soil plant comparisons as established.
Strandberg et al. (2025) compared capture probe, metabarcoding, and shotgun sequencing approaches for vegetation characterisation and found that different methods captured different components of diversity, underlining the value of multi-approach design.
3.3 Air vs forensic approaches for terrestrial vertebrates
Newton et al. (2026) conducted the most rigorous multi-substrate comparison for terrestrial vertebrates, comparing airborne eDNA against spider webs, soil, leaf swabs, and other passive substrates. Airborne eDNA and spider webs together outperformed other substrates for total species richness detected, while soil was less effective for highly mobile species such as birds and bats. Berard et al. (2025) confirmed that no single substrate dominated across all taxonomic groups and environments, reinforcing the complementarity principle.
4. DNA persistence: a key difference between substrates
How long DNA persists in each medium fundamentally shapes what a sample represents temporally.
| Substrate | Typical DNA persistence | What the sample represents |
|---|---|---|
| Flowing water | Hours to days | Recent past (hours upstream) |
| Lake/pond water | Days to weeks | Recent occupancy of water body |
| Soil (surface) | Days to months | Past season to year |
| Lake sediment | Decades to millennia | Historical ecosystem archive |
| Air (active sampling) | Minutes to hours | Current atmospheric signal |
| Air (passive dust trap) | Days to weeks | Recent period of deposition |
| Archived air filters | Decades (if stored correctly) | Historical snapshot at collection date |
The short persistence of DNA in air (where UV degradation and dry deposition act rapidly) is both a limitation (cannot archive biodiversity retrospectively unless filters are deliberately preserved) and an advantage (detections are current). The discovery that archived radionuclide filters retain recoverable DNA for decades under appropriate storage conditions (Sullivan et al. 2025) demonstrates that the historic analysis window can be dramatically extended with intentional archival protocols.
5. Spatial scale: the fundamental contrast
The most important practical difference between the three substrates is their spatial scale of operation.
A single water eDNA sample from a river represents species present upstream within a specific hydrological catchment — typically a few kilometres of channel, if we neglect contamination sources. A soil core or swab represents a patch of a few square metres to, at most, a few hundred square metres. An air sample represents a landscape footprint of several square kilometres to several tens of kilometres, depending on atmospheric conditions and sampler volume.
This scaling difference has direct implications for monitoring programme design:
The corollary is that airborne eDNA is most powerful as a landscape-level biodiversity indicator rather than a site-level species inventory tool. It is the method of choice when the question is "what is the overall state of biodiversity across this region?" rather than "does this specific pond contain great crested newts?"
6. Combining substrates: the future of eDNA monitoring
The scientific community increasingly recognises that no single substrate is optimal across all questions and taxa. Altermatt et al. (2025) argue for integrated multi-substrate eDNA monitoring frameworks that use water, soil and air sampling in complementary roles within national monitoring networks.
A practical integrated framework might look like this:
This multi-substrate architecture provides complementary spatial scales, temporal resolutions, and taxonomic coverage, while using existing infrastructure wherever possible.
7. Caveats and open questions
Direct substrate comparisons are rare. Most studies use a single substrate. The few direct comparisons that exist (Newton et al. 2026; Berard et al. 2025) are at single sites and cannot be generalised to all habitat types or taxa. Systematic multi-substrate comparison studies across diverse ecosystems are an urgent research priority.
Cost-effectiveness benchmarking is immature. Tournayre et al. (2025) showed airborne eDNA has low marginal cost when using existing infrastructure, but no study has yet conducted a rigorous cost-per-species-detected comparison across substrates for the same sites, questions and taxa.
The cross-medium transfer question is advancing but not resolved. Ip et al. (2025a, 2025b) showed water-to-air eDNA transfer for both single species (salmon qPCR) and community-level vertebrate assemblages (metabarcoding). The community-level data show that abundant species track reliably across media, while rare taxa drop out — implying that cross-medium transfer is real but detection-limited for low-abundance organisms. How this generalises across water body types (lentic vs lotic, marine vs freshwater) and environmental conditions remains to be established. An analogous question applies to air-to-soil transfer: DNA deposited from the atmosphere onto soil surfaces could in principle be recovered from soil samples, particularly after rain events that wash particulate matter into the upper soil layer. This pathway has received almost no direct study but would imply that soil eDNA near intensive land use areas (e.g. arable fields, roads, high-traffic natural sites) may contain atmospheric DNA contributions that are not of strictly local origin. This is a research gap with direct relevance to soil eDNA monitoring programme design.
References
- Altermatt F et al. (2025). Utilizing aquatic environmental DNA to address global biodiversity targets. Nature Reviews Biodiversity. https://doi.org/10.1038/s44185-025-00063-x
- Berard J et al. (2025). Spider webs, soil or leaf swabs to detect environmental DNA from terrestrial vertebrates. Molecular Ecology Resources. https://doi.org/10.1111/1755-0998.70037
- Bowers RM et al. (2011). Sources of bacteria in outdoor air across cities in the midwestern United States. Applied and Environmental Microbiology 77:6350–6356. https://doi.org/10.1128/AEM.05498-11
- Deiner K et al. (2017). Environmental DNA metabarcoding: Transforming how we survey animal and plant communities. Molecular Ecology 26:5872–5895. https://doi.org/10.1111/mec.14350
- Jane SF et al. (2015). Distance, flow regime, and PCR inhibition control detection probabilities of spiny crayfish environmental DNA. Molecular Ecology Resources 15:562–574. https://doi.org/10.1111/1755-0998.12285
- Johnson M & Barnes MA (2024). Macrobial airborne environmental DNA analysis: A review. Molecular Ecology Resources 24:e13998. https://doi.org/10.1111/1755-0998.13998
- Lynggaard C et al. (2024). Airborne environmental DNA captures terrestrial vertebrate diversity in nature. Molecular Ecology Resources 24:e13840. https://doi.org/10.1111/1755-0998.13840
- Newton JP et al. (2026). Airborne DNA and spider webs outperform other eDNA sources for monitoring terrestrial vertebrates. Molecular Ecology Resources. https://doi.org/10.1111/1755-0998.70067
- Ip YCA et al. (2025a). Passive air sampling detects environmental DNA transfer from water into air. Scientific Reports. https://doi.org/10.1038/s41598-025-26293-6
- Ip YCA et al. (2025b). Vertebrate biodiversity via eDNA at the air-water interface. iScience. https://doi.org/10.1016/j.isci.2026.115682
- Shogren AJ et al. (2017). Controls on eDNA movement in streams: Transport, retention, and resuspension. Scientific Reports 7:5065. https://doi.org/10.1038/s41598-017-05223-1
- Strandberg NA et al. (2025). Capture probe, metabarcoding, or shotgun sequencing: which reflects local vegetation best? bioRxiv. https://doi.org/10.1101/2025.10.06.680654
- Sullivan AR et al. (2025). Airborne eDNA captures three decades of ecosystem biodiversity. Nature Communications 16:11281. https://doi.org/10.1038/s41467-025-67676-7
- Thomsen PF & Willerslev E (2015). Environmental DNA — An emerging tool in conservation for monitoring past and present biodiversity. Biological Conservation 183:4–18. https://doi.org/10.1016/j.biocon.2014.11.019
- Tournayre O et al. (2025). First national survey of terrestrial biodiversity using airborne eDNA. Scientific Reports. https://doi.org/10.1038/s41598-025-03650-z
- Willerslev E et al. (2003). Diverse plant and animal genetic records from Holocene and Pleistocene sediments. Science 300:791–795. https://doi.org/10.1126/science.1084114