Short Answer: Weather sits at the intersection of the two forces that govern what an air sample contains: transport and degradation. Wind moves DNA-laden particles from their sources towards samplers — but also dilutes them and extends the catchment area. Rain removes particles from the air column. UV radiation and heat degrade DNA, shortening the window of detectability. Temperature and humidity shape both organism activity (and therefore DNA release rates) and DNA stability in transit. The result is that every air sample is the product of a balance between these competing forces, and understanding that balance is essential for interpreting what a detection means and designing protocols that are temporally and spatially representative.
1. Weather as the frame for transport and degradation
Airborne eDNA exists in a dynamic system. Unlike a water sample — where the DNA concentration reflects a relatively stable pool of recent shedding — an air sample captures a snapshot of a highly variable atmospheric flux. The composition and concentration of biological particles in air changes by the hour in response to meteorological conditions.
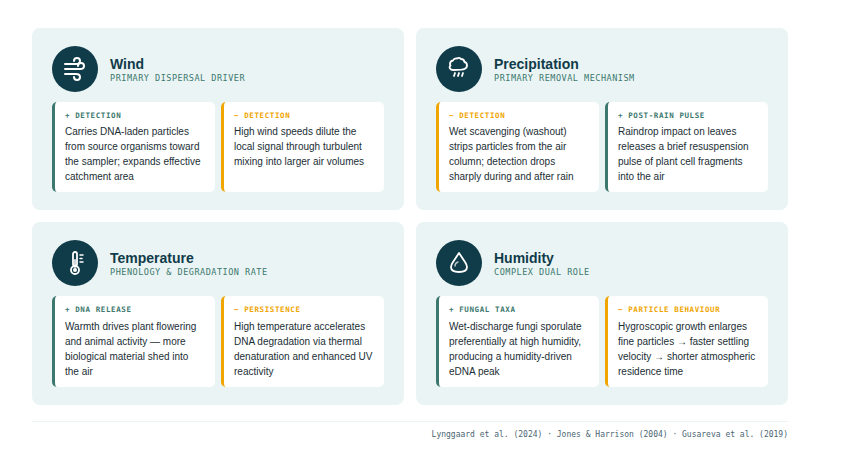
The key conceptual frame is the tension between two competing processes. Transport moves DNA-carrying particles from source organisms towards the sampler; weather conditions that favour transport (wind, turbulence, dry air) tend to increase the diversity and quantity of signal detected. Degradation destroys the DNA in those particles; conditions that accelerate degradation (UV radiation, heat, chemical oxidants in the atmosphere) reduce the window over which a detection remains possible. These two forces pull in partially opposite directions: a long, windy, sunny day transports DNA further but also degrades it faster. A cool, overcast, calm period preserves DNA longer but moves it less. The signal a sampler captures is always the net result of both.
A third factor sits alongside these: organism activity. Animals move, shed DNA, and release faecal aerosols more intensively during warm, active periods. Plants flower and release pollen on temperature cues. Fungi discharge spores at specific humidity thresholds. The same meteorological variables that govern transport and degradation also govern how much DNA enters the air in the first place — amplifying or dampening the signal at source before it even begins to travel.
Aerobiology has documented weather-driven variability in airborne pollen and spore concentrations for decades (Hirst 1952). Applying this understanding to the broader macrobial eDNA context is the work of the current generation of researchers.
2. Wind: the primary dispersal mechanism
2.1 Wind speed and horizontal transport
Wind is the dominant force determining how far airborne eDNA travels and how quickly it arrives at a sampler. At its most basic, higher wind speeds carry particles further and faster from their source. This has two opposing effects on detection:
Positive effect: Wind transports DNA from source organisms to samplers, particularly over distances of hundreds of metres to kilometres. Lynggaard et al. (2024) observed that PM10 particle concentrations varied across sampling events and speculated that wind speed may influence the transport of airborne eDNA.
Negative effect (dilution): Very high winds increase turbulent mixing, diluting the local DNA signal by mixing it into a larger air volume. For detecting locally present species, this can reduce detection probability despite increasing particle transport.
The net effect on any given study depends on scale: at landscape scale, moderate winds generally improve detection by distributing DNA from distributed source populations. At site scale (a few hundred metres), calmer conditions may actually favour detection of species present in the immediate vicinity by allowing local DNA to accumulate.
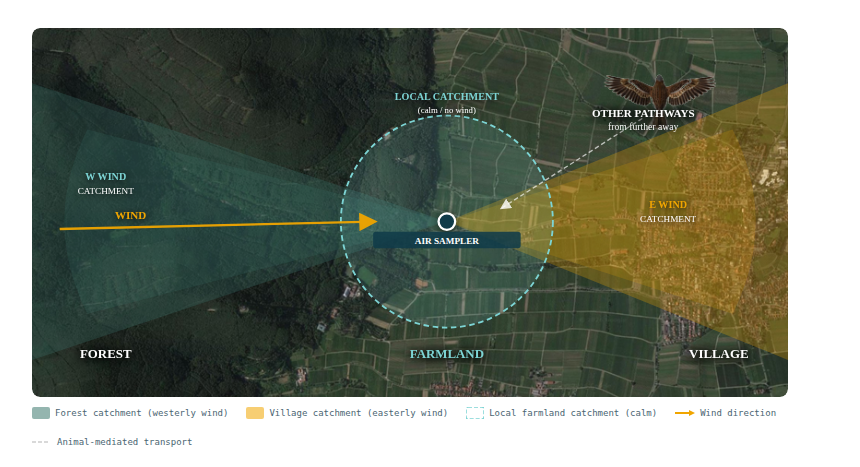
2.2 Wind direction and catchment shape
Wind direction determines the source region from which a sampler draws its DNA signal. On any given day, the sampler primarily captures organisms from the upwind sector. Sullivan et al. (2025) demonstrated this elegantly using HYSPLIT back-trajectory analysis, showing that the effective catchment of their Swedish radionuclide station was elongated in the direction of prevailing winds and shifted seasonally with large-scale atmospheric circulation.
This directional effect has important practical implications: the same sampler in the same location will have different effective catchment areas and therefore different species compositions depending on seasonal wind patterns. Multi-season monitoring is essential for obtaining a representative picture of the local biodiversity community. And if a specific location has to be monitored multiple sample locations may have to be selected to accomodate changing wind directions.
2.3 Turbulence, vertical mixing, and nocturnal transport
Turbulence — irregular, chaotic air motion — has two effects. In the surface layer (0–50 m above ground), turbulence mixes biological particles into the airstream and keeps them suspended. In the planetary boundary layer (~0–1,500 m), daytime convective turbulence lifts particles high into the atmosphere, increasing transport potential. At night, turbulence collapses and particles accumulate in a shallow layer near the ground, producing higher near-surface concentrations that can benefit low-mounted samplers.
Two additional nocturnal phenomena are particularly relevant for eDNA monitoring in flat terrain such as the Netherlands. The nocturnal low-level jet is a wind speed maximum that forms at the top of the nocturnal stable boundary layer, typically between 100–500 m altitude. While surface winds are calm at night, this elevated jet can transport biological particles horizontally over tens of kilometres — meaning near-ground samplers may capture material that arrived overnight from a different source region than daytime samples from the same location. Fumigation events occur during the morning transition when the boundary layer grows rapidly as solar heating resumes: biological material that was transported overnight at altitude within the residual layer is brought down to the surface as the mixed layer deepens. Fumigation produces a brief pulse of increased airborne particle concentrations at ground level, often around mid-morning. Both phenomena are well-documented in aerobiology for pollen transport in the Dutch landscape (Stull 1988, An Introduction to Boundary Layer Meteorology) and add to the complexity of interpreting early-morning or overnight airborne eDNA samples.
This diurnal pattern in atmospheric stability means that sampling time of day can influence what is detected. Also seasonal patterns in the 34-year Sullivan et al. (2025) archive varied with atmospheric conditions. Gusareva et al. (2019) documented strong diel cycles in tropical airborne microbial communities, with community composition differing markedly between day and night — with substantially higher concentrations observed at night, when turbulent daytime mixing subsides and near-surface accumulation occurs.
3. Precipitation: the primary removal mechanism
3.1 Wet deposition and rain scavenging
Rain is the most efficient removal mechanism for biological particles from the atmosphere. Falling raindrops collide with and capture airborne particles (a process called wet scavenging or washout), depositing them onto surfaces and effectively removing them from the air column. Wet scavenging (washout) can substantially reduce airborne biological particle concentrations during and after rain events, but raindrop can also enhance production or release particles in the air (Jones & Harrison 2004).
Lynggaard et al. (2024) found that vertebrate DNA detection rates in their Danish forest study decreased following rainfall events, hinting towards rain-driven removal of DNA-laden particles from the sampled air. This is one of the most consistent and reproducible weather effects documented in the airborne eDNA literature.
The practical implication is that samples collected immediately after heavy rain will reflect a depleted atmospheric signal — and may underestimate local biodiversity. Post-rain negative detections should not be interpreted as evidence of species absence.
3.2 Post-rain recovery and soil resuspension
After rain stops and conditions dry, biological particles are resuspended from wet vegetation and soil surfaces into the air, potentially producing a brief post-rain pulse in airborne DNA concentration. Hanson et al. (2024) found that storm events were associated with elevated plant leaf fragment DNA in the air, attributing this to the mechanical action of rain impacting vegetation and releasing cellular material. High wind events during or after storms may amplify this effect.
3.3 Fog and wet scavenging efficiency
Fog and dew represent wet deposition without active rainfall. Fog droplets settle slowly onto surfaces, capturing airborne particles. However, the efficiency of wet scavenging — whether by fog, drizzle, or rain — is strongly size-dependent, with important implications for the practical advice often given to eDNA practitioners.
Below-cloud rain scavenging operates primarily through two mechanisms: inertial impaction (efficient for coarse particles >10 µm, including most pollen grains) and Brownian diffusion (efficient for very fine particles <0.1 µm). Particles in the intermediate range of approximately 1–10 µm — which includes many vertebrate eDNA carriers such as skin cell fragments, hair fragments, and sub-pollen particles — are the least efficiently scavenged by either mechanism. This means that moderate rainfall reduces the atmospheric eDNA signal substantially for large particles (pollen, large spores) but may have a less dramatic effect on the small-particle vertebrate eDNA fraction than is commonly assumed.
The practical implication is that the standard recommendation to “avoid sampling immediately after rain” applies most strongly to studies targeting large-particle taxa such as pollen or large fungal spores. For vertebrate metabarcoding using markers that amplify DNA from small cell fragments and aerosols, the post-rain depletion effect may be real but shorter-lived, and the specific reduction in signal will depend on the particle size distribution of the relevant DNA carriers — something that has not been systematically quantified in the airborne eDNA context.
4. Temperature: effects on DNA release and degradation
4.1 Biological release mechanisms
Temperature affects the rate at which organisms shed biological material. For plants, temperature is the primary driver of phenological transitions: flowering onset, leaf expansion, and leaf senescence are all temperature-dependent. The start of the pollen season in temperate Europe can be predicted with considerable accuracy using accumulated temperature sums (Linkosalo et al. 2010; Sofiev et al. 2013). These temperature-driven phenological patterns are clearly reflected in airborne eDNA signals: Sullivan et al. (2025) found that plant eukaryotes showed single seasonal peaks tightly coupled to temperature-driven phenology.
For animals, temperature affects activity levels, reproductive behaviour, and movement rates — all of which influence DNA shedding. During cold winter months, reduced animal activity and movement likely reduces the total DNA flux entering the atmosphere, though this has not been directly measured.
4.2 Temperature and DNA persistence
Higher temperatures accelerate DNA degradation through thermal denaturation and enhanced enzymatic activity. UV radiation, more intense in summer and at lower latitudes, is the primary outdoor driver. The practical implication is that summer detections may represent a shorter temporal window than winter detections — the DNA was present more recently. Controlled quantification of these degradation rates under outdoor airborne conditions remains limited; most data come from aquatic eDNA systems and cannot be applied directly to the atmospheric context.
5. Humidity: a complex dual role
5.1 Effects on particle behaviour
Relative humidity affects both biological particle release and atmospheric particle behaviour. Many fungal species release spores preferentially during periods of specific humidity — either dry conditions (for dry-discharge species) or wet conditions following rainfall (for wet-discharge species). This produces characteristic humidity-dependent patterns in fungal eDNA that have been documented in aerobiology for decades.
For animal-derived DNA, high humidity can cause hygroscopic growth of fine particles — they absorb water vapour and increase in size, which increases settling velocity and reduces atmospheric residence time. Conversely, very dry conditions may increase the friability of leaf and skin material, potentially increasing fragmentation and aerosolisation of DNA-carrying particles.
5.2 DNA stability and humidity
Humidity has a complex relationship with DNA stability. At intermediate humidity levels, DNA associated with protective particles (cells, pollen grains) is relatively stable. At very high humidity, water can facilitate enzymatic degradation if associated organisms remain metabolically active. At very low humidity, physical damage through desiccation can fragment DNA.
Although Harnpicharnchai et al. (2023) sampled within a subway they found seasonal variation in active airborne eDNA collection related to particulate matter (PM), temperature, and relative humidity, recommending that these variables be recorded alongside every sampling event as covariates for interpretation.
6. Seasonal patterns: integrating meteorological effects
The combined effects of all meteorological variables produce systematic seasonal patterns in airborne eDNA signals that reflect both biological (phenological) and physical (meteorological) drivers:
7. Practical implications for study design and interpretation
- Always record weather data alongside eDNA sampling.Wind speed and direction, precipitation (including events before and during sampling), temperature, and relative humidity should be logged as covariates. Without this, meteorological confounding cannot be accounted for in analysis.
- Avoid sampling immediately after heavy rain.Although not fully certain it would be advisable to maintain a minimum of 6–12 hours post-rain before active sampling, where feasible. Continuous passive samplers are less affected by this concern as they integrate over the full weather cycle.
- Multi-season sampling is essential for community characterisation.No single season captures all taxa. A monitoring programme that samples only during summer will miss autumn fungal diversity, winter bacterial signals, and spring plant phenology.
- Interpret wind-associated detections cautiously.High-wind detections of species that seem unexpected for the local habitat may reflect regional transport from distant sources. Back-trajectory analysis should be considered for anomalous detections.
- Continuous monitoring outperforms spot samples.Systems that collect air continuously (like air quality monitoring stations) automatically integrate across weather variability and provide far more robust community characterisation than any individual spot sample.
References
- Lynggaard C et al. (2024). Airborne environmental DNA captures terrestrial vertebrate diversity in nature. Molecular Ecology Resources 24:e13840. https://doi.org/10.1111/1755-0998.13840
- Sullivan AR et al. (2025). Airborne eDNA captures three decades of ecosystem biodiversity. Nature Communications 16:11281. https://doi.org/10.1038/s41467-025-67676-7
- Johnson M & Barnes MA (2024). Macrobial airborne environmental DNA analysis: A review. Molecular Ecology Resources 24:e13998. https://doi.org/10.1111/1755-0998.13998
- Jones AM & Harrison RM (2004). The effects of meteorological factors on atmospheric bioaerosol concentrations. Science of the Total Environment 326:151–180. https://doi.org/10.1016/j.scitotenv.2003.11.021
- Sofiev M et al. (2013). A numerical model of birch pollen emission and dispersion in the atmosphere. International Journal of Biometeorology 57:45–58. https://doi.org/10.1007/s00484-012-0532-z
- Harnpicharnchai P et al. (2023). Temporal dynamics in diversity and composition of the air mycobiome and dominant allergenic fungi in the subway environment. Environmental DNA 5:1265–1288. https://doi.org/10.1002/edn3.449
- Hanson M et al. (2024). Storms facilitate airborne DNA from leaf fragments outside the main tree pollen season. Aerobiologia 40(3):415–423. https://doi.org/10.1007/s10453-024-09826-w
- Gusareva ES et al. (2019). Microbial communities in the tropical air ecosystem follow a precise diel cycle. PNAS 116:23299–23308. https://doi.org/10.1073/pnas.1908493116
- Niu M et al. (2021). Influence of rainfall on fungal aerobiota in the urban atmosphere over Tianjin, China: A case study. Atmospheric Environment: X 12:100137. https://doi.org/10.1016/j.aeaoa.2021.100137
- Linkosalo T et al. (2010). A double-threshold temperature sum model for predicting the flowering duration of Betula pendula. Tree Physiology 30:904–916. https://doi.org/10.1093/treephys/tpq025
- Strickler KM et al. (2015). Quantifying effects of UV-B, temperature, and pH on eDNA degradation. Biological Conservation 183:85–92. https://doi.org/10.1016/j.biocon.2014.11.038
- Abrego N et al. (2024). Airborne DNA reveals predictable spatial and seasonal dynamics of fungi. Nature 631(8022):835–842. https://doi.org/10.1038/s41586-024-07658-9
- Hirst JM (1952). An automatic volumetric spore trap. Annals of Applied Biology 39:257–265. https://doi.org/10.1111/j.1744-7348.1952.tb00904.x
- Stull RB (1988). An Introduction to Boundary Layer Meteorology. Kluwer Academic Publishers, Dordrecht. ISBN 978-9027727688.
- Lin R et al. (2025). Robust passive sampling of airborne environmental DNA. Methods in Ecology and Evolution. https://doi.org/10.1111/2041-210X.70102